Current issue
Archive
Manuscripts accepted
About the Journal
Editorial office
Editorial board
Section Editors
Abstracting and indexing
Subscription
Contact
Ethical standards and procedures
Most read articles
Instructions for authors
Article Processing Charge (APC)
Regulations of paying article processing charge (APC)
IMMUNOLOGY / BASIC RESEARCH
Molecular cloning, expression, purification and in silico epitope prediction of cobalamin-independent methionine synthase (Mor a 2), as a novel allergen from Morus alba pollen
1
Department of Molecular Biology and Genetics, Institute of Graduate Studies in Sciences, Istanbul University, Istanbul, Turkey
2
Department of Molecular Biology and Genetics, Faculty of Science, Istanbul University, Istanbul, Turkey
3
Division of Immunology and Allergic Diseases, Department of Internal Medicine, Istanbul Faculty of Medicine, Istanbul University, Istanbul, Turkey
4
Center for Research and Practice in Biotechnology and Genetic Engineering (BIYOGEM), Istanbul University, Istanbul, Turkey
Submission date: 2020-10-10
Final revision date: 2021-04-26
Acceptance date: 2021-05-01
Online publication date: 2021-05-12
Corresponding author
Yunus Aksüt
Department of Molecular Biology and Genetics, Institute of Graduate Studies in Sciences, Istanbul University, Istanbul, Turkey
Department of Molecular Biology and Genetics, Institute of Graduate Studies in Sciences, Istanbul University, Istanbul, Turkey
KEYWORDS
Morus albapollen allergyrecombinant allergenMor a 2cobalamin-independent methionine synthase (MetE)Schizosaccharomyces pombeB- and T-cell epitopes
TOPICS
ABSTRACT
Introduction:
Morus alba (white mulberry) pollen is an aero-allergen source that can trigger allergic diseases. Cobalamin-independent methionine synthase (MetE) in M. alba pollen has been proved to be one of the major allergens for some patients living in Istanbul (Turkey). The aim of the present study was to carry out recombinant production and identification of MetE (Mor a 2), a novel allergen from M. alba pollen. The IgE binding reactivity of rMor a 2 produced for the first time was evaluated and some structural features were investigated by in silico methods to better understand its immunogenicity.
Material and methods:
The gene encoding Mor a 2 was cloned in fission yeast, Schizosaccharomyces pombe ura4-D18h- strain, using the pSLF1073 vector. This is the first report of the production of recombinant pollen allergen in S. pombe. After the purification, immunoreactivity of rMor a 2 was confirmed by immunoblotting using sera of a patient allergic to M. alba pollen. Moreover, B-cell epitopes of rMor a 2 were predicted using various bioinformatic tools, namely Bioinformatics Predicted Antigenic Peptides, BepiPred 2.0 and Immune Epitope Database, whereas T-cell epitopes were estimated using NetMHCIIpan-3.2 and NetMHCII 2.3 servers.
Results:
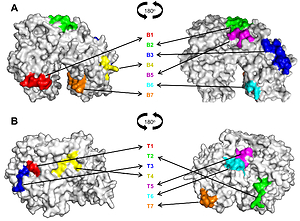
The immunoblotting analysis yielded 11 of 11 positive reactions to rMor a 2. In silico predictions exerted seven B-cell epitopes (22-33, 384-394, 407-423, 547-553, 571-577, 671-678, 736-741) and seven T-cell epitopes (54-62, 161-170, 197-205, 347-358, 622-630, 657-665, 756-764).
Conclusions:
These findings may support the use of rMor a 2 in the diagnosis and treatment of allergic diseases associated with M. alba and/or MetE.
Morus alba (white mulberry) pollen is an aero-allergen source that can trigger allergic diseases. Cobalamin-independent methionine synthase (MetE) in M. alba pollen has been proved to be one of the major allergens for some patients living in Istanbul (Turkey). The aim of the present study was to carry out recombinant production and identification of MetE (Mor a 2), a novel allergen from M. alba pollen. The IgE binding reactivity of rMor a 2 produced for the first time was evaluated and some structural features were investigated by in silico methods to better understand its immunogenicity.
Material and methods:
The gene encoding Mor a 2 was cloned in fission yeast, Schizosaccharomyces pombe ura4-D18h- strain, using the pSLF1073 vector. This is the first report of the production of recombinant pollen allergen in S. pombe. After the purification, immunoreactivity of rMor a 2 was confirmed by immunoblotting using sera of a patient allergic to M. alba pollen. Moreover, B-cell epitopes of rMor a 2 were predicted using various bioinformatic tools, namely Bioinformatics Predicted Antigenic Peptides, BepiPred 2.0 and Immune Epitope Database, whereas T-cell epitopes were estimated using NetMHCIIpan-3.2 and NetMHCII 2.3 servers.
Results:
The immunoblotting analysis yielded 11 of 11 positive reactions to rMor a 2. In silico predictions exerted seven B-cell epitopes (22-33, 384-394, 407-423, 547-553, 571-577, 671-678, 736-741) and seven T-cell epitopes (54-62, 161-170, 197-205, 347-358, 622-630, 657-665, 756-764).
Conclusions:
These findings may support the use of rMor a 2 in the diagnosis and treatment of allergic diseases associated with M. alba and/or MetE.
REFERENCES (46)
1.
Puc M. Characterisation of pollen allergens. Ann Agric Environ Med 2003; 10: 143-50.
2.
Radauer C, Breiteneder H. Pollen allergens are restricted to few protein families and show distinct patterns of species distribution. Allergy Clin Immunol 2006; 117: 141-7.
3.
Alessandri C, Zennaro D, Zaffiro A, Mari A. Molecular allergology approach to allergic diseases in the paediatric age. Ital J Pediatr 2009; 35: 29.
4.
Muñoz FJ, Delgado J, Palma J, Giménez MJ, Monteseirín FJ, Conde J. Airborne contact urticaria due to mulberry (Morus alba) pollen. Contact Dermatitis 1995; 32: 61.
5.
Navarro AM, Orta JC, Sánchez MC, Delgado J, Barber D, Lombardero M. Primary sensitization to Morus alba.
7.
Singh BP, Singh AB, Nair PK, Gangal SV. Survey of airborne pollen and fungal spores at Dehra Dun, India. Ann Allergy 1987; 59: 229-34.
8.
Sneller MR, Hayes HD, Pinnas JL. Pollen changes during five decades of urbanization in Tucson, Arizona. Ann.
10.
Ciardiello MA, Palazzo P, Bernardi ML, et al. Biochemical, immunological and clinical characterization of a cross-reactive nonspecific lipid transfer protein 1 from mulberry. Allergy 2010; 65: 597-605.
11.
Micheal S, Wangorsch A, Wolfheimer S. Immunoglobulin E reactivity and allergenic potency of Morus papyrifera (paper mulberry) pollen. J Investig Allergol Clin Immunol 2013; 23: 168-75.
12.
Choi JH, Sim JK, Oh JY, et al. An IgE-mediated allergic reaction caused by mulberry fruit. Allergy Asthma Immunol Res 2015; 7: 195-8.
13.
Aslam MS, Khalid T, Gull I, et al. Identification of major allergens of paper mulberry (Broussonetia papyrifera) pollens and purification of novel 40 kDa allergen protein. Curr Allergy Clın Im 2015; 28: 36-41.
14.
Çetereisi D, Karlioglu N, Gelincik A, et al. Proteomic identification of allergenic proteins of Morus alba L. Asian Pac J Allergy Immunol 2019; 37: 205-11.
15.
Ferrer JL, Ravanel S, Robert M, Dumas R. Crystal structures of cobalamin-independent methionine synthase complexed with zinc, homocysteine, and methyltetrahydrofolate. Biol Chem 2004; 279: 44235-8.
16.
Chardin H, Mayer C, Senechal H, Tepfer M, Desvaux FX, Peltre G. Characterization of high-molecular-mass allergens in oilseed rape pollen. Int Arch Allergy Immunol 2001; 125: 128-34.
17.
Assarehzadegan MA, Sankian M, Jabbari F, Tehrani M, Falak R, Varasteh A. Identification of methionine synthase (Sal k 3), as a novel allergen of Salsola kali pollen. Mol Biol Rep 2011; 38: 65-73.
18.
Schmidt M, Hoffman DR. Expression systems for production of recombinant allergens. Int Arch Allergy.
20.
Cui Y, Yu L, Teng F, et al. Expression of recombinant allergen, Der f 1, Der f 2 and Der f 4 using baculovirus-insect cell systems. Arch Med Sci 2018; 14: 1348-54.
21.
Valenta R, Linhart B, Swoboda I, Niederberger V. Recombinant allergens for allergen-specific immunotherapy: 10 years anniversary of immunotherapy with recombinant allergens. Allergy 2011; 66: 775-83.
22.
Jutel M, Kosowska A, Smolinska S. Allergen immunotherapy: past, present, and future. Allergy Asthma Immunol Res 2016; 8: 191-7.
23.
Ciprandi G, De Amici M, Murdaca G, et al. Serum IL-4 as a marker of immunological response to sublingual immunotherapy. J Biol Regul Homeost Agents 2008; 22: 117-23.
24.
Ciprandi G, De Amici M, Negrini S, Marseglia G, Tosca MA. TGF-beta and IL-17 serum levels and specific immunotherapy. Int Immunopharmacol 2009; 9: 1247-9.
25.
Gregan J, Rabitsch PK, Rumpf C, Novatchkova M, Schleiffer A, Nasmyth K. High-throughput knockout screen in fission yeast. Nat Protoc 2006; 1: 2457-64.
26.
Gasteiger E, Hoogland C, Gattiker A, et al. Protein identification and analysis tools on the ExPASy server. In: Walker JM (ed.). The Proteomics Protocols Handbook. Humana Press, New Jersey 2005; 571-607.
27.
Jones DT. Protein secondary structure prediction based on position-specific scoring matrices. J Mol Biol 1999; 292: 95-202.
28.
Petersen B, Petersen TN, Andersen P, Nielsen M, Lundegaard C. A generic method for assignment of reliability scores applied to solvent accessibility predictions. BMC Struct Biol 2009; 9: 51.
29.
Arnold K, Bordoli L, Kopp J, Schewde T. The SWISS-MODEL workspace: a web-based environment for protein structure homology modelling. Bioinformatics 2006; 22: 195-201.
30.
Laskowski RA, MacArthur MW, Moss DS, Thornton MJ. PROCHECK: a program to check the stereochemical quality of protein structures. J Appl Cryst 1993; 26: 283-91.
31.
Colovos C, Yeates TO. Verification of protein structures: patterns of nonbonded atomic interactions. Protein Sci 1993; 2: 1511-9.
32.
Bowie JU, Lüthy R, Eisenberg D. A method to identify protein sequences that fold into a known three-dimensional structure. Science 1991; 253: 164-70.
33.
Benkert P, Tosatto SC, Schomburg D. QMEAN: A comprehensive scoring function for model quality assessment. Proteins 2008; 71: 261-77.
34.
Wiederstein M, Sippl MJ. ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 2007; 35: W407-10.
35.
Jespersen MC, Peters B, Nielsen M. BepiPred-2.0: improving sequence-based B-cell epitope prediction using conformational epitopes. Nucleic Acids Res 2017; 45: W24-9.
36.
Yang X, Yu X. An introduction to epitope prediction methods and software. Rev Med Virol 2009; 19: 77-96.
37.
Wang DW, Ni WW, Zhou YJ, et al. Expression, purification and epitope analysis of Pla a 2 allergen from Platanus acerifolia pollen. Mol Med Rep 2018; 17: 394-9.
38.
Kurowski M, Majkowska-Wojciechowska B, Wardzyńska A, Kowalski ML. Associations of allergic sensitization and clinical phenotypes with innate immune response genes polymorphisms are modified by house dust mite allergen exposure. Arch Med Sci 2011; 7: 1029-36.
39.
Oeo-Santos C, Mas S, Benedé S, et al. A recombinant isoform of the Ole e 7 olive pollen allergen assembled by de novo mass spectrometry retains the allergenic ability of the natural allergen. J Proteomics 2018; 187: 39-46.
40.
Paddock CD, McKerrow JH, Hansell E, Foreman KW, Hsieh I, Marshall N. Identification, cloning, and recombinant expression of procalin, a major triatomine allergen. J Immunol 2001; 167: 2694-9.
41.
Morín M, Asturias JA, Domínguez A. Expression of Alt a 1 allergen from Alternaria alternata in the yeast Yarrowia lipolytica. FEMS Microbiol Lett 2012; 333: 121-8.
42.
De Silva C, Dhanapala P, King S, Timothy Doran T, Tang M, Suphioglu C. Immunological comparison of native and recombinant Hen’s egg yolk allergen, chicken serum albumin (Gal d 5), produced in Kluveromyces lactis. Nutrients 2018; 10: 757.
43.
Sasaki M, Kumagai H, Takegawa K, Tohda H. Characterization of genome-reduced fission yeast strains. Nucleic Acids Res 2013; 41: 5382-99.
44.
Zhang Z, Cai Z, Hou Y, et al. Enhanced sensitivity of capture IgE ELISA based on a recombinant Der f 1/2 fusion protein for the detection of IgE antibodies targeting house dust mite allergens. Mol Med Rep 2019; 19: 3497-504.
45.
Eberlein B. Basophil activation as marker of clinically relevant allergy and therapy outcome. Front Immunol 2020; 11: 1815.
46.
Nair S, Kukreja N, Singh BP, Arora N. Identification of B cell epitopes of alcohol dehydrogenase allergen of Curvularia lunata. PloS one 2011; 6: e20020.
We process personal data collected when visiting the website. The function of obtaining information about users and their behavior is carried out by voluntarily entered information in forms and saving cookies in end devices. Data, including cookies, are used to provide services, improve the user experience and to analyze the traffic in accordance with the Privacy policy. Data are also collected and processed by Google Analytics tool (more).
You can change cookies settings in your browser. Restricted use of cookies in the browser configuration may affect some functionalities of the website.
You can change cookies settings in your browser. Restricted use of cookies in the browser configuration may affect some functionalities of the website.