Current issue
Archive
Manuscripts accepted
About the Journal
Editorial office
Editorial board
Section Editors
Abstracting and indexing
Subscription
Contact
Ethical standards and procedures
Most read articles
Instructions for authors
Article Processing Charge (APC)
Regulations of paying article processing charge (APC)
NEPHROLOGY / RESEARCH PAPER
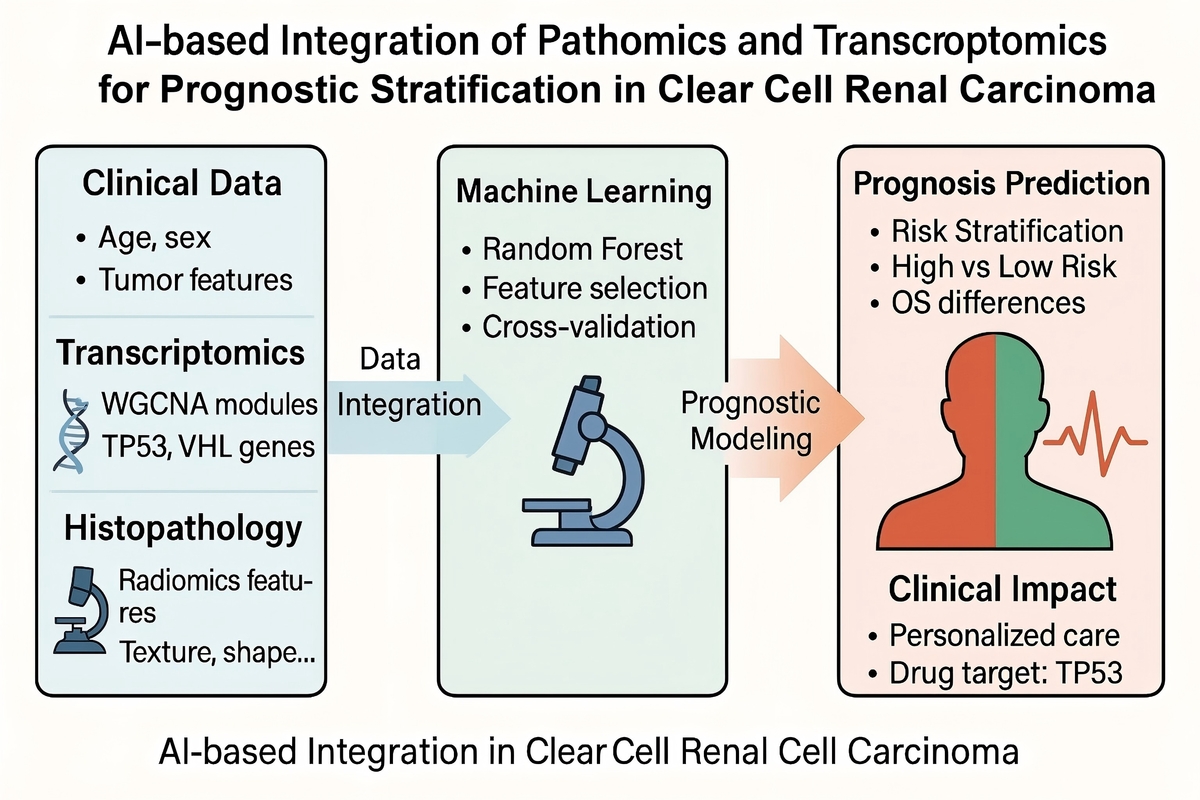
Application of AI-based Machine Learning Integrating Pathomics and Transcriptomics for Diagnosis and Prognostic Prediction in Clear Cell Renal Cell Carcinoma
1
The Fourth Affiliated Hospital of China Medical University, China
Submission date: 2025-07-19
Final revision date: 2025-12-12
Acceptance date: 2026-02-05
Online publication date: 2026-05-01
KEYWORDS
Artificial IntelligenceDeep LearningTranscriptomicsClear Cell Renal Cell CarcinomaPathological OmicsPrognostic Prediction
TOPICS
ABSTRACT
Introduction:
Clear cell renal cell carcinoma (ccRCC) is the most common and aggressive subtype of renal carcinoma, characterized by high molecular heterogeneity and variable clinical outcomes. Conventional prognostic models often lack the resolution to adequately stratify patient risk, underscoring the need for integrative, data-driven approaches.
Material and methods:
We developed a novel diagnostic and prognostic model for ccRCC using an artificial intelligence-based machine learning approach (random forest algorithm) integrating pathomics and transcriptomic data. Whole-slide histopathological images and transcriptomic data were obtained from the TCGA-KIRC cohort. Radiomic features were extracted from histological slides, and gene modules related to tumor progression were identified using Weighted Gene Co-Expression Network Analysis (WGCNA). These modalities, together with clinical variables, were incorporated into a custom machine learning architecture to predict patient survival risk.
Results:
The integrative model demonstrated strong predictive performance, effectively stratifying patients into high- and low-risk groups with significant differences in overall survival (p < 0.001). Functional enrichment analysis revealed that prognostic gene modules were associated with immune regulation, angiogenesis, and cell cycle pathways, highlighting their relevance in ccRCC pathogenesis.
Conclusions:
This study presents a novel, AI-driven framework that combines multi-omics and imaging data to improve prognostic accuracy in ccRCC. The model offers potential utility in clinical decision-making and personalized treatment strategies, and may serve as a foundation for future precision oncology applications.
Clear cell renal cell carcinoma (ccRCC) is the most common and aggressive subtype of renal carcinoma, characterized by high molecular heterogeneity and variable clinical outcomes. Conventional prognostic models often lack the resolution to adequately stratify patient risk, underscoring the need for integrative, data-driven approaches.
Material and methods:
We developed a novel diagnostic and prognostic model for ccRCC using an artificial intelligence-based machine learning approach (random forest algorithm) integrating pathomics and transcriptomic data. Whole-slide histopathological images and transcriptomic data were obtained from the TCGA-KIRC cohort. Radiomic features were extracted from histological slides, and gene modules related to tumor progression were identified using Weighted Gene Co-Expression Network Analysis (WGCNA). These modalities, together with clinical variables, were incorporated into a custom machine learning architecture to predict patient survival risk.
Results:
The integrative model demonstrated strong predictive performance, effectively stratifying patients into high- and low-risk groups with significant differences in overall survival (p < 0.001). Functional enrichment analysis revealed that prognostic gene modules were associated with immune regulation, angiogenesis, and cell cycle pathways, highlighting their relevance in ccRCC pathogenesis.
Conclusions:
This study presents a novel, AI-driven framework that combines multi-omics and imaging data to improve prognostic accuracy in ccRCC. The model offers potential utility in clinical decision-making and personalized treatment strategies, and may serve as a foundation for future precision oncology applications.
We process personal data collected when visiting the website. The function of obtaining information about users and their behavior is carried out by voluntarily entered information in forms and saving cookies in end devices. Data, including cookies, are used to provide services, improve the user experience and to analyze the traffic in accordance with the Privacy policy. Data are also collected and processed by Google Analytics tool (more).
You can change cookies settings in your browser. Restricted use of cookies in the browser configuration may affect some functionalities of the website.
You can change cookies settings in your browser. Restricted use of cookies in the browser configuration may affect some functionalities of the website.