Therapeutic modification of atherogenic lipoproteins by statins [1], ezetimibe [2] and Proprotein convertase subtilisin/kexin type 9 (PCSK9) inhibition [3] is an effective strategy to reduce the risk of cardiovascular disease (CVD), principally by lowering non-HDL-cholesterol [4]. PCSK9 regulates LDL-cholesterol (LDL-C) through hepatic expression of LDL receptors. PCSK9 inhibitors are licensed as LDL-C lowering agents with excellent efficacy and evidence of cardiovascular benefits: the FOURIER trial [3] reported that a monoclonal antibody to PCSK9 that lowered LDL-C by 59% (~56 mg/dl) led to a 15% reduction in the risk of a composite of cardiovascular death, myocardial infarction, stroke, hospitalization for unstable angina or coronary revascularization in patients with established atherosclerotic vascular disease. However, early phase 3 studies showed a potential excess of neurocognitive adverse effects [5], leading the US FDA to instruct pharmaceutical companies to assess potential neurocognitive side effects of PCSK9 inhibitors. While, reassuringly, no excess risk was noted in the FOURIER trial [3] or its substudy with detailed cognitive measures [6], these studies cannot fully exclude potential adverse effects from longer-term use of PCSK9 inhibitors. An orthogonal approach to obtain reliable information is to exploit genetic variants that mimic pharmacological inhibition of PCSK9. Naturally occurring variation in the gene encoding a drug target can be used to gauge insight on long-term effects of therapeutic modification [7]. So-called drug-target Mendelian randomization [8] exploits the characteristics of the genotype for the reliable estimation of both intended and unintended consequences of therapeutic modification of a drug target, as previously demonstrated [9–12].
Methods
We used four measures of cognitive ability from UK Biobank (UKB) data: two from baseline (2006-2010): fluid reasoning measured in 160,130 individuals (with genetic data, prior to exclusions listed below) and reaction time in 482,187 as they showed good intra-participant longitudinal reliability in n = 19,999 participants [13], and two measured in 2014–2015 via the internet: trail making test (TMT) A (processing speed) in 100,587 and B (speed plus executive function) in 100,610, and digit symbol coding (executive function) in 115,933. TMT and reaction time scores were log-transformed due to a positive skew. All measures were standardized to Z-scores.
Six independent single nucleotide polymorphisms (SNPs) (R2 < 0.15) in PCSK9 (referent allele frequencies rs2479394 A = 0.72; rs11206510 C = 0.19; rs2479409 A = 0.65; rs10888897 T = 0.39; rs7552841 C = 0.63; rs562556 G = 0.18); orientated so that the effect allele associated with a lower LDL-C were used as genetic variants to proxy therapeutic inhibition of PCSK9. SNPs were selected from the paper by Ference et al. in NEJM [13]. In the paper by Ference et al. [14], seven PCSK9 SNPs were used, however two were found to be moderately correlated (rs2479409 and rs2149041; R2 = 0.37). We therefore removed the SNP with the weaker association with LDL-C (rs2149041) as defined by the p-value [14] leaving six SNPs (with pair-wise LD R2 < 0.15) described above. The LDL-C association of each of the six PCSK9 SNPs from the Global Lipids Genetics Consortium [15] was used to construct a weighted PCSK9 allele score.
Ethical approval
This secondary-data analysis study was conducted under the generic approval from the NHS National Research Ethics Service (approval letter dated 17th June 2011, ref. 11/NW/0382). Written informed consent was obtained from all participants in the study (consent for research, by UK Biobank).
Statistical analysis
Linear regression analyses used the four cognition traits as dependent variables, the weighted PCSK9 score as the independent variable, adjusted for age, sex, GWAS array, and 10 principal components (as provided by UKB). To mimic pharmacological modification of PCSK9, we report results of the PCSK9 allele score scaled to the 50 mg/dl lower LDL-C achieved in FOURIER. We compared estimates of the PCSK9 allele score with cognition traits to those from APOE e4 dosage (excluding rare APOE e2/e4), APOE e4/e4 vs. e3/e3 homozygosity; current vs. never smoking, and 5-years of increased cross-sectional age with cognitive traits. As a further positive control, we tested the association of the PCSK9 allele score with the risk of CHD in UK Biobank (defined as self-reported physician-diagnosed myocardial infarction and angina).
We excluded participants with non-white British ancestry, self-report vs. genetic sex mismatch, putative sex chromosomal aneuploidy, excess heterozygosity, and missingness rate > 0.1. We removed one random participant in cases where two individuals were first cousins or closer. Stata v14 and PLINK v1.90 were used for analyses.
Data availability statement
UK Biobank is an open access resource available to verified researchers upon application (http://www.ukbiobank.ac.uk/). Analysis syntax is available upon request.
Results
The PCSK9 allele score scaled to a 50 mg/dl lower LDL-C was associated with a lower risk of CHD in UKB (comprising 15,284 cases of myocardial infarction and angina in 338,852 individuals; OR = 0.73; 95% CI: 0.60–0.90, p = 0.003).
We next investigated the associations of six variants in PCSK9 scaled to 50 mg/dl lower LDL-C in 109,870 individuals with measures of fluid reasoning, 337,348 with reaction time, 73,044 with processing speed (TMT A), 73,063 with processing speed plus executive function (TMT B), and 82,012 with digit symbol coding (executive function) (Figure 1).
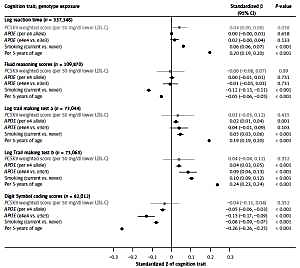
Figure 1
Association of a PCSK9 allele score (in gray) scaled to 50 mg/dl lower LDL-cholesterol and other selected exposures (in black) with cognitive traits in UK Biobank
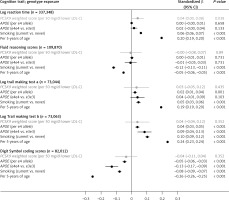
Orientated to a 50 mg/dl lower LDL-C (i.e. mimicking pharmacological inhibition of PCSK9), the PCSK9 allele score was nominally associated with log reaction time (0.04 SDs higher log reaction time; 95% CI: 0.002–0.079; p = 0.038). For fluid reasoning, the scaled PCSK9 allele score had wide 95% CI (–0.08, 0.07) that included the estimates for the association of 5 years additional age (–0.05 SDs, 95% CI: –0.06, –0.05). Similar patterns were identified for all other cognition traits, meaning that despite the large sample size, the imprecision around the estimates obtained from the PCSK9 allele score meant that we could not exclude a similar magnitude of effect of genetic inhibition of PCSK9 to that seen with the positive controls, including APOE e4 or smoking status for any of the individual cognition traits (Figure 1). Notably, point estimates for the association of the PCSK9 allele score and all cognitive ability end-points were on the harmful side of unity. In sensitivity analyses, removal of participants that self-reported a neurological condition (~5% of the dataset) [13] did not alter the findings, nor did substituting rs2479409 for rs2149041 in the PCSK9 allele score.
Discussion
In this large-scale analysis of individuals from the general population, we used naturally occurring genetic variants in PCSK9 to gauge insight into the effect of lifelong lowering of LDL-C through inhibition of PCSK9 and its association with cognition abilities. Using available data in UKB, we were not able to provide definitive evidence on the relationship of PCSK9 genetic variants with cognition traits. While this may have arisen due to lack of power, we note firstly that we were able to show associations of the PCSK9 allele score with the risk of prevalent self-reported CHD, and secondly robust associations of conventional risk factors (e.g. age and smoking) and genetic variants (e.g. APOE e4) with the cognitive traits. These observations suggest validity of the PCSK9 genetic score as an instrument, and sufficient statistical power for detecting association with relevant traits.
Our findings add to previous studies of genetically-estimated PCSK9 and cognition: Schmidt et al. [16] reported no significant association in nine cohorts, three of which indexed cognition with a screening tool (Mini-mental state exam) rather than normative-range tests as per here; while Rao et al. reported no effect on a single cognitive test (Trail-making; processing speed and executive function) [17]. The cognitive tests used here were novel, brief and bespoke to UK Biobank baseline assessment [13], and therefore examinations of cognitive assessments which are based on more traditional and validated tasks, may be informative. Such analyses may also test for potential age- and sex-specific associations.
While the imprecision makes it challenging to draw firm conclusions about an effect (or lack thereof) of life-long LDL-cholesterol lowering by PCSK9 inhibition on cognition, our data highlight the need for additional large-scale genetic analyses. In parallel, continued pharmacovigilance is potentially needed for patients currently treated with PCSK9 inhibitors.