Introduction
Bladder cancer is a highly recurrent malignancy [1, 2]. Although cytotoxic chemotherapeutic agents such as gemcitabine (GEM) initially work well for most patients, the majority of patients treated with GEM progressively generate resistance after multiple rounds of therapy, eventually leading to tumor recurrence [3, 4]. Chemotherapies are usually administered in cycles to kill asynchronously proliferating cancer cells, at the same time allowing the restoration of cells in normal tissue [5, 6]. However, this therapeutic approach also provides an opportunity for residual cancer cells to survive [3]. Results from recent studies suggest that a small subpopulation of cancer cells with stem cell-like properties has a survival advantage in response to chemotherapy and may be responsible for tumor recurrence [7, 8]. This subpopulation of cancer cells is also known as cancer stem cell-like cells (CSCs) or tumor-initiating cells (TIC) since they retain the ability to self-renew and repopulate the entire tumor mass [9]. However, the molecular mechanisms underlying the GEM-induced enrichment of CSCs in urothelial bladder cancer cells (UBCs) remain unclear, which hinders the application of therapies specifically targeting CSCs.
Circular RNAs (circRNAs) are novel non-coding RNAs with end loop structure and are widely found in mammalian cells [10, 11]. The main characteristics of circRNAs are their tissue or cell specificity and high stability [12]. In general, circRNAs can fulfill their functions through various pathways. For example, circRNAs compete as miRNA sponges for binding to microRNAs (miRNAs), which consequently regulate gene transcription [13]. In addition, increasing evidence of circRNA dysregulation in a variety of diseases, such as triple-negative breast cancer, gastric cancer, and pancreatic ductal adenocarcinoma, suggests that circRNAs are involved in cancer development [14]. Studies have shown the presence of a large number of circRNA transcripts in tumor cells, highlighting their great potential as biomarkers for cancer diagnosis and treatment [14]. Among them, cricRNA_103809 has been reported to act as a pro-oncogenic factor in colorectal and lung cancers [15, 16]. However, its role and molecular mechanism in bladder cancer are still unknown.
In this study, we aimed to explore the expression and function of cricRNA_103809 in bladder cancer and identified the miR-516a/FBXL18 axis as a downstream regulatory mechanism.
Material and methods
Cell culture
Bladder cancer cell lines EJ, T24T, UMUC3, RT4, 5637 and the urothelial cell line UROtsa were obtained from Procell (Wuhan, China). EJ and 5637 cells were cultured in RPMI-1640 medium, T24T and RT4 cells in McCoy 5A medium, and UMUC3 cells in MEM medium containing 10% fetal bovine serum (FBS). Gemcitabine-resistant cells were incubated in culture medium that contains a low dose of gemcitabine (0.5 µM). All cells were incubated in a 37°C atmosphere with 5% CO2.
Cell transfection
The circRNA-103809, shRNA, miR-516a mimics and inhibitors, and FBXL18 overexpression vectors were synthesized by RiboBio (Guangzhou, China) and were transfected into cells as per the protocol of Lipofectamine 2000 (Invitrogen, USA).
Fluorescence in situ hybridization (FISH) assay
The localization of circRNA-103809 was analyzed using an In Situ Hybridization Kit following the manufacturer’s protocol. In brief, the Cy3-labeled circRNA-103809 probe was incubated with cells at 37°C in an incubator overnight. The images of cells were captured using a confocal microscope (Carl Zeiss, Germany).
qPCR assay
Total RNA was extracted from cultured cells and tissues using Trizol reagent (Thermo, USA). Then cDNA was synthesized by using a PrimeScript RT Reagent Kit (Takara, Japan). Gene expression was measured using a TB Green Premix Ex Taq kit (Takara, Japan) and calculated according to the 2–ΔΔCt method. The actin level was adopted as an internal control.
Cell Counting Kit 8 (CCK-8)
To measure cell proliferation, 5,000 cancer cells from each group were seeded into each well of a 96-well plate. After culture for 24, 48, 72, and 96 h, cells were reacted with 10 µl of CCK-8 solution for 2 h. Then absorbance at 450 nm was measured using a microplate reader (Tanon, China), and cell viability was calculated.
Transwell assay
Cell migration and invasion was measured by a Transwell experiment. Cells were digested and suspended in culture medium containing 0.1% FBS, then added into the upper chamber of a 12-well Transwell plate. Culture medium containing 10% FBS was added to the lower chamber. After culture for 24 h, the upper chamber was collected, and cells were stained with 0.1% crystal violet. Images were captured with a microscope (Carl Zeiss, Germany).
Sphere formation assay
T24 and 5637 cancer cells were suspended as single cells and seeded into a 96-well ultra-low attachment plate (Corning, USA) with 1000 cells in each well. Cell culture medium was composed of DMEM/F-12 supplemented with 20 ng/ml basic fibroblast growth factor (bFGF, Invitrogen, USA), 20 ng/ml epidermal growth factor (EGF, Invitrogen, USA), and B27 reagent (Invitrogen, USA). The cells were cultured at 37°C incubator for 14 days, then spheres with a diameter larger than 50 µm were counted.
Western blotting assay
After treatment, cells were lysed with RIPA buffer (CWBIO, China) and centrifuged. The supernatant was collected, and protein concentration was measured using a BCA assay kit (Beyotime, China). A total of 35 µg of protein was separated by SDS-PAGE and transferred to polyvinylidene difluoride (PVDF) membranes, followed by blocking with 5% skimmed milk at room temperature. The membranes were probed with E-cadherin, N-cadherin, vimentin, Snail1, ALDH1, Gil-1, OCT4, Smo, FBXL18, and β-actin overnight at 4°C. The next day, HRP-conjugated secondary antibodies were applied to incubate with the protein bands. Then the protein bands were visualized after reaction with ECL reagent (Millipore, Germany).
RNA pulldown
The biotinylated wild type (WT) and mutated (MUT) miR-516a probes were synthesized using a Biotin RNA Labeling Kit (Beyotime, China). T24 and 5637 cells were collected and homogenized with co-IP buffer. After centrifuging, the supernatant was collected and hatched with the biotin-labeled miRNAs for one night at 4°C, followed by incubation with streptavidin-coated magnetic beads (Invitrogen, USA) for 60 min at room temperature. The captured circRNAs were eluted from the beads and analyzed by qPCR assay.
Luciferase reporter gene assay
The sequences of wild type or mutated circ_103809 and 3′UTR of FBXL18 were cloned into pmirGLO plasmid and were co-transfected with miR-516a mimics into the bladder cancer cells using Lipofectamine 2000 (Invitrogen, USA) following the manufacturer’s protocol. The transfected cells were incubated for 24 h and lysed to measure luciferase activities using the dual-luciferase reporter assay system (Promega, USA) as per the manufacturer’s introduction.
Xenograft tumor model
BALB/c nude mice aged 4 weeks were bought from Vital River Laboratory and acclimated in a standard environment for 1 week. Bladder cancer cells were suspended in serum free medium at a density of 5 × 106/100 µl, and 100 µl of cell suspension was inoculated into the right flank of each mouse. Each group contained 5 mice. The width and length of tumors were measured every 3 days using a slide caliper, and tumor volume was calculated as follows: volume (mm3) = 0.5 × length × width2. When the tumor size reached 100 mm3, the treatment with siRNA and miRNA inhibitors (10 nmol/20 g body weight) began and was conducted every 3 days. All mice were sacrificed by pentobarbital calcium after cell injection for 30 days, and tumors were collected. Animal experiments in this research were performed under the guidelines authorized by the Animal Ethics Committee of the First Affiliated Hospital of Wenzhou Medical University. The animal experiments were approved by the Animal Ethical Committee of the First Affiliated Hospital of Wenzhou Medical University (2022071308).
Statistical analysis
All data in this study were shown as mean ± standard deviation (SD) and analyzed using GraphPad Prism 8.0 software. The difference between two or more parametric variables was analyzed by Student’s t-test or one-way analysis of variance (ANOVA), respectively. The difference between nonparametric variables was determined by Mann-Whitney U test. p < 0.05 was set as the criterion of statistical significance.
Results
CircRNA_103809 knockdown suppresses proliferation, migration, and invasion of bladder cancer cells in vitro
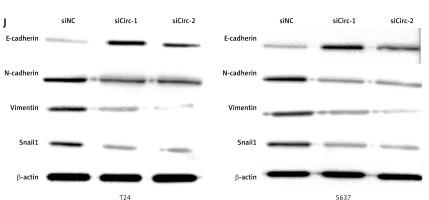
To determine the effects of circRNA_103809 on bladder cancer cells, we first analyzed its level in normal and malignant cells. As shown in Figure 1 A, the level of circRNA_103809 was notably higher in bladder cancer cell lines compared with the urothelial cell line UROtsa, among which T24 and 5637 showed the highest level and were selected for subsequent experiments. The level of circRNA_103809 was not obviously changed within 24 h in physical condition (Figure 1 B) and RNase R treatment (Figure 1 C), whereas the level of linear RNA ZFR was obviously decreased, indicating the stability of circRNA_103809 as circRNA. Results from the FISH experiment showed the cytoplasm localization of circRNA_103809, suggesting it may function in cytoplasm at the post-transcriptional level. We next conducted knockdown and overexpression of circRNA_103809 using siRNAs (Figure 1 E) and overexpression vectors (Figure 1 F). We observed significantly repressed growth (Figure 1 G), migration, and invasion (Figures 1 H and I) of bladder cancer cells under knockdown of circRNA_103809. Additionally, the protein level of E-cadherin was elevated, and N-cadherin, vimentin, and Snail levels were reduced under treatment with siRNA targeting circRNA_103809 (Figure 1 J). These data indicated that circRNA_103809 suppressed proliferation, migration, and invasion of bladder cancer cells in vitro.
Figure 1
CircRNA_103809 silencing suppresses proliferation, migration, and invasion of bladder cancer cells in vitro. A – Levels of circRNA_103809 in urothelial cell line UROtsa and bladder cancer cells. B – Levels of circRNA_103809 and linear RNA ZFR within 24 h were measured by qPCR. C – Level of circRNA_103809 under treatment of RNase R was measured by qPCR. D – Intracellular localization of circRNA_103809 was detected by FISH assay. E – Level of circRNA 103809 in bladder cancer cells after transfection with sicircRNA_103809. F – Level of circRNA_103809 in bladder cancer cells after transfection with circRNA_103809 overexpression vectors. G – Growth of bladder cancer cells after transfection with sicircRNA_103809. H, I – Migration and invasion of bladder cancer cells after transfection with sicircRNA_103809. J – Protein levels of E-cadherin, N-cadherin, vimentin, and Snail in bladder cancer cells were measured by western blotting assay. *p < 0.05, **p < 0.01, ***p < 0.001 *p < 0.05, **p < 0.01, ***p < 0.001
CircRNA_103809 affects stemness and gemcitabine resistance of bladder cancer cells
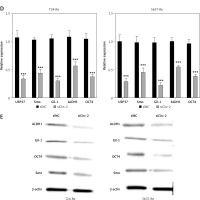
Next, we explored the effects of circRNA_103809 on drug resistance of bladder cancer cells. We observed a significantly elevated level of circRNA_103809 in gemcitabine-resistant bladder cancer cells compared with the parental cells (Figure 2 A). Knockdown of circRNA_103809 suppressed the viability of drug resistant cancer cells under gemcitabine treatment (Figure 2 B). Also, the sphere formation ability of gemcitabine-resistant bladder cancer cells was notably suppressed by sicircRNA_103809 (Figure 2 C), along with significantly decreased RNA and protein levels of cancer cell stemness markers USP37, Smo, Gil-1, ALDH1, and OCT4 (Figures 2 D and E).
Figure 2
CircRNA_103809 affects stemness and gemcitabine resistance of bladder cancer cells. A – Level of cicRNA_103809 in gemcitabine-resistant and parental bladder cancer cells. B – Viability of gemcitabine-resistant and parental bladder cancer cells under treatment with gemcitabine. C – Sphere formation ability of gemcitabine-resistant bladder cancer cells under transfection with sicircRNA_103809. D, E – RNA and protein levels of cancer cell stemness markers USP37, Smo, Gil-1, ALDH1, and OCT4. ***p < 0.001
Si-CircRNA_103809 represses in vivo growth of bladder cancer cells
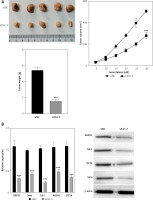
We further confirmed the effects of circRNA_103809 on in vivo growth of bladder cancer cells. We established a xenograft tumor model using T24 cells transfected with siNC or sicircRNA_103809. As shown in Figure 3 A, the tumor size, growth rate, and tumor weight were significantly suppressed in the sicircRNA_103809 group compared with the siNC group. Moreover, the RNA and protein levels of stemness markers ALDH1, Gil-1, OCT4, and Smo in tumor tissues were notably decreased by knockdown of circRNA_103809 (Figure 3 B).
Figure 3
Si-CircRNA_103809 represses the in vivo growth of bladder cancer cells. Xenograft tumor model was established by using T24 cells. A – Tumor size, tumor growth curve, and tumor weight were recorded. B – RNA and protein levels of stemness markers ALDH1, Gil-1, OCT4, and Smo in tumors were measured. ***p < 0.001
CircRNA_103809 promotes expression of FBXL18 by sponging of miR-516a
Subsequently, we investigated the regulatory mechanisms underlying the effects of circRNA_103809. We analyzed the potential interaction between circRNA_103809 with miR-516a. Transfection with miR-516a effectively upregulated the level of miR-516a (Figure 4 A) and repressed the activity of wild type circRNA_103809 luciferase reporter gene vectors rather than the mutated vectors (Figure 4 B). Results from RNA pulldown indicated that circRNA_103809 could directly interact with miR-516a (Figure 4 C). Furthermore, luciferase of wild type FBXL18 reporter gene vectors was significantly reduced by miR-516a (Figure 4 D). Additionally, the protein level of FBXL18 was notably suppressed by miR-516a (Figure 4 E) and sicircRNA_103809 (Figure 4 F), which was recovered by miR-516a inhibitors (Figure 4 F). These data indicated that circRNA_103809 interacted with miR-516a to modulate FBXL18 expression.
Figure 4
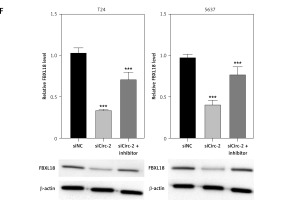
CircRNA_103809 promotes expression of FBXL18 by sponging of miR-516a. A – T24 cells and 5637 cells were transfected with miR-516a, then level of miR-516a was measured by qPCR. B – Luciferase activity of wild type and mutated circRNA_103809 reporter gene vectors under treatment with miR-516a. C – RNA pulldown assay to measure interaction between biotin-labeled wild type or mutated miR-516a. D – Luciferase activity of wild type and mutated FBXL18 3′UTR under treatment with miR-516a. E – Protein level of FBXL18 in T24 cells and 5637 cells was measured by western blotting assay. F – Protein level of FBXL18 in T24 cells and 5637 cells was measured by western blotting assay. *p < 0.05, **p < 0.01, ***p < 0.001
CircRNA_103809 targets the miR-516a/FBXL18 signaling pathway to modulate bladder cancer cell phenotypes
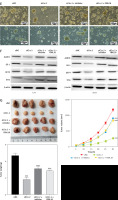
We next performed cellular and animal experiments to verify the circRNA_103809/miR-516a/FBXL18 regulatory axis using sicirc_103809, miR-516a inhibitors, and FBXL18 overexpression vectors. The results from CCK-8 and Transwell assay showed that knockdown of circRNA_103809 significantly suppressed the growth, migration and invasion of bladder cancer cells, whereas inhibition of miR-516a and overexpression of FBXL18 could reverse these inhibitory effects (Figures 5 A–C), along with repressed expression of E-cadherin and elevation of N-cadherin, vimentin, and Snail (Figure 5 D). The sicircRNA_103809-suppressed sphere formation (Figure 5 E) and expression of ALDH1, Gil-1, OCT4, and Smo were also recovered by miR-516a inhibitors and FBXL18 overexpression vectors (Figure 5 F). Consistently, the in vivo anti-tumor effects of sicirc_103809 were also repressed by miR-516a inhibitors and FBXL18 overexpression vectors (Figure 5 G).
Figure 5
CircRNA_103809 targets the miR-516a/FBXL18 signaling pathway to modulate bladder cancer cell phenotypes. A–F – T24 and 5637 cells were transfected with sicircRNA_103809, miR-516a inhibitors, and FBXL18 overexpression vectors. Then (A) cell viability. (B and C) cell migration and invasion, (D) protein levels of cell migration and invasion biomarkers, (E) sphere formation ability, and (F) protein expression of stemness markers were measured. G – A xenograft model was established and treated with sicircRNA_103809, miR-516a inhibitors, and FBXL18 overexpression vectors. Then tumor size, tumor growth curve, and tumor weight were measured. (E) sphere formation ability, and (F) protein expression of stemness markers were measured. G – A xenograft model was established and treated with sicircRNA_103809, miR-516a inhibitors, and FBXL18 overexpression vectors. Then tumor size, tumor growth curve, and tumor weight were measured. ***p < 0.001
Discussion
In this study, we investigated the anti-tumor effects of circRNA_103809 on bladder cancer and found that knockdown of circRNA_103809 could significantly suppress the in vitro and in vivo growth of bladder cancer cells, reduced cell migration and invasion, increased sensitivity to gemcitabine, and reduced cancer cell stemness. Further exploration of molecular mechanisms revealed that circRNA_103809 interacted with miR-516a to modulate the expression of FBXL18.
In recent decades, circRNAs have been widely reported as critical regulators in various physical and pathological processes [17–19]. Mechanistically, circRNAs could directly interact with proteins or miRNAs [20, 21]. On the other hand, miRNA targets the specific mRNA sequence to inhibit the translation of mRNA or induce its degradation, which consequently suppresses expression of the target gene [22, 23]. Increasing evidence has shown that circRNAs participate in regulation of cell growth, differentiation, migration, metabolism, etc. [12]. Targeting circRNAs is a potential strategy for cancer therapy [24, 25]. CircRNA_103809 is reported to be a potential tumor promoter in several cancer types. For example, analysis of clinical samples of hepatocellular carcinoma demonstrated a dysregulated level of circRNA_103809, and circRNA_103809 could upregulate the protein level of fibroblast growth factor receptor 1 (FGFR1) via sponging miR-377-3p to modulate cell proliferation, migration, and invasion [26]. CircRNA_103809 expression is elevated in gastric cancer tissues and cancer cell lines. Meanwhile, knockdown of circRNA_103809 remarkably reduced the migration and invasive ability of gastric cancer cells [27]. Similarly, our findings also showed that knockdown of circRNA_103809 effectively suppressed the growth, migration, and invasion of bladder cancer cells. We also found that gemcitabine-resistant bladder cancer cells showed decreased viability and impaired stemness under knockdown of circRNA_103809. A previous study obtained similar results, finding that circRNA_103809 knockdown enhanced the sensitivity of breast cancer cells to gemcitabine, which is correlated with miR-516a-5p-regulated F-box and leucine-rich repeat protein 18 (FBXL18) expression [28]. FBXL18 is an E3 ubiquitin ligase that is frequently reported to participate in the progression of multiple cancers. FBXL18 depletion suppressed the proliferation and cell growth and induced apoptosis of glioma cells via inhibiting AKT activity and phosphorylation of FOXO3 [29]. In hepatocellular carcinoma cells, FBXL18 promotes ubiquitination of RPS15A and increases the translocation of SMAD3 to the nucleus to promote cancer cell growth [30]. Similarly, we identified FBXL18 as a mediator of circRNA_103809-regulated bladder cancer growth. While our preclinical findings are promising, we must emphasize the need for rigorous clinical validation to assess whether targeting circRNA_103809 will yield consistent effects in human patients. The translational gap between preclinical and clinical studies in oncology is well documented, and the challenges associated with RNA-based therapies – including specificity, delivery, and immunogenicity – are fundamental barriers that must be addressed. Furthermore, alternative explanations for the dysregulation of circRNA_103809 must be explored. It is possible that its expression changes could be secondary to other oncogenic processes, rather than indicating a primary role in cancer progression. To strengthen our conclusions, systematic analysis of circRNA_103809 expression across diverse cancer types, stages, and patient cohorts is essential. Future studies should leverage advanced techniques such as single-cell RNA sequencing and spatial transcriptomics, and incorporate patient-derived samples to correlate circRNA_103809 expression with clinical outcomes. We also plan to discuss technological innovations, such as CRISPR-based RNA editing and novel delivery systems, to address current limitations in circRNA-targeted therapies. By acknowledging these challenges and the speculative nature of some claims, we aim to provide a more nuanced and scientifically rigorous analysis of circRNA_103809’s role in cancer biology.
Limitations: 1) While we suggest that circRNA_103809 may have broader relevance as a cancer biomarker, it is crucial to recognize the highly context-dependent nature of circRNA biology. CircRNAs typically participate in specific regulatory networks that can vary significantly based on the tumor microenvironment and molecular subtype. Recent studies indicate that circRNAs can exhibit contrasting roles – both oncogenic and tumor-suppressive – depending on the cellular context. Therefore, without robust cross-cancer comparisons, our assertion regarding circRNA_103809 as a universal biomarker remains to be thoroughly evaluated. 2) Although our findings from in vitro and in vivo models provide valuable insights, these models do not fully capture the complexities of human cancers. For instance, murine models often fail to replicate the heterogeneity and immune interactions present in human tumors. Thus, caution is warranted in extrapolating our preclinical findings directly to clinical outcomes. 3) Our optimistic outlook on circRNA-targeted therapies does not sufficiently address significant challenges, including specificity – achieving selective targeting of circRNA_103809 without affecting other RNA molecules remains a substantial hurdle. Off-target effects could result in unintended gene dysregulation. Additionally, effective and safe delivery of RNA therapies to tumors poses ongoing challenges. Existing methods, such as lipid nanoparticles and viral vectors, face obstacles such as immune clearance and limited tumor penetration. Furthermore, circRNA-based therapies may provoke innate immune responses, complicating their therapeutic application. 4) CircRNA expression can vary significantly with disease stage and treatment status. Failing to account for these temporal dynamics could lead to overgeneralized conclusions about circRNA_103809’s role across different cancer stages. 5) We recognize the importance of interpatient variability in circRNA expression, which may influence the utility of circRNA_103809 as a biomarker. Factors such as genetic background and prior treatments could impact its expression and function, underscoring the need for large-scale clinical studies. 6) Our claim regarding circRNA_103809 as a universal biomarker requires further empirical support. Given the documented heterogeneity of circRNA expression across various cancers, it is essential to avoid overly ambitious generalizations about its role.
In conclusion, we identified potential promoting effects of circRNA_103809 during bladder cancer cell growth, while depletion of circRNA_103809 repressed cell growth, migration, invasion, and stemness, and enhanced sensitivity to gemcitabine. Mechanistic investigation further revealed the circRNA_103809/miR-516a/FBXL18 regulatory axis in bladder cancer. We have identified circRNA_103809 as a promising therapeutic target for bladder cancer therapy.