Introduction
HIV/AIDS (acquired immunodeficiency syndrome) has become a major global public health problem since it was first reported in 1981 [1]. Approximately 37.9 million people are currently living with HIV-1 and there are 1.7 million newly infected patients worldwide each year (www.unaids.org). Because an effective HIV vaccine has not been developed yet, the use of anti-HIV drugs has become the main method for controlling the HIV epidemic. Although antiretroviral therapy, especially highly active antiretroviral therapy (HARRT), has greatly reduced HIV/AIDS-related mortality and increased the quality of life of many patients, the current therapy has failed to cure HIV infection [2–4]. Furthermore, the toxic side effects [5], drug resistance, and high mutation rate of HIV often lead to treatment failure [6, 7]. Therefore, it is expected that novel anti-HIV will be discovered, thus promoting the development of anti-HIV therapy.
In generally, HIV-1 replication occurs in a six-step life cycle, including binding and fusion, reverse transcription, integration, transcription, assembly, and budding [8]. So far, three key viral enzymes in HIV-1 replication, including reverse transcriptase (RT) [9], integrase (IN) [10], and protease (PR), are the main targets for the development of anti-HIV-1 drugs [11, 12]. HIV RT transforms the single-stranded viral RNA genome to double-stranded DNA [12]. HIV DNA is processed and integrated into the host cell genome by viral IN [13]. PR plays an important role in the mature of the virus [14]. Because the human host does not have the similar proteins as these viral enzymes, HIV-1 RT, IN, and PR are crucial targets in study of new anti-HIV drugs due to their importance and uniqueness in viral replication [15]. Antiretroviral drugs acting on different viral targets have proved successful in HAART [16]. Therefore, researchers prefer to find multi-target drugs with multi-activities against different steps of HIV-1 replication [16].
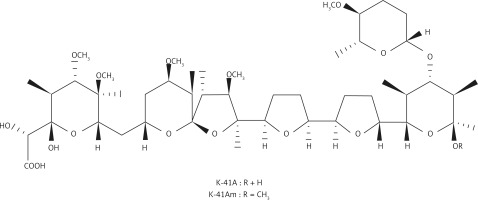
In our screening of novel anti-HIV-1 compounds, we found that two polyether compounds, antibiotics K-41A (formerly designated as K-41) and K-41Am (Figure 1), had relatively strong anti-HIV activity. K-41Am, named by us, is a new compound. These two compounds were isolated from the fermentation broth of Streptomyces sp. SCSIO 01680, which was isolated from sediment samples in the northern South China Sea. According to WESTLEY classification, polyethers are classified into four groups by structure: monovalent polyether, monovalent monoglycoside polyether, divalent polyether, and divalent pyrrole ether [17]. K-41A and K-41Am belong to monoglycoside polyethers.20 K-41A is a known antibiotic with activities against gram-positive bacteria and coccidia [18, 19], and is a promising lead compound for new oral antimalarial drugs [20]. K-41Am is a methylated derivative of K-41A. The only difference between the two compounds is the R substituents [18, 21]. The molecular structures of K-41A and K-41Am are shown in Figure 1. K-41A and K-41Am attach a terminal (F) ring [22]. This terminal (F) ring is rare and was only previously found in K-41A, K-41B [22], and CP-96,797 [21]. In this study, we characterised the anti-HIV-1 activity of K-41A and K-41Am as well as the mechanisms involved.
Material and methods
Bacterial strain, fermentation, isolation, and structure identification of antibiotics K-41A and K-41Am
The antibiotics K-41A and K-41Am used in this study were isolated from fermentation broth of strain SCSIO 01680. The strain was isolated from a sediment sample in the northern South China Sea and was identified as Streptomyces sp. SCSIO 01680 based on morphology and 16S rRNA sequence analysis. Modified RA medium (1.0% glucose, 2.0% soluble starch, 1.0% malt meal, 1.0% maltose, 0.5% corn steep liquor, 3.0% sea salt pH 7.2–7.4) was used for large-scale fermentation. For isolation of K-41 and K-41Am, 15 l of fermentation broth was extracted with butanone, and the organic extract was evaporated to dryness to yield a residue. The residue was suffered from bioassay-guided fractionation against Micrococcus luteus over normal phase silica gel column chromatography and preparative thin-layer chromatography (TLC) to yield K-41A (20.5 mg) and K-41Am (23.0 mg). The structures of antibiotics K-41A and K-41Am were characterised by high-resolution mass spectrometer (HRMS), 1H, 13C, and 2D nuclear magnetic resonance (NMR) spectroscopic analysis as well as comparisons with those reported in the literature [18, 19].
Chemical reagents
Tested compounds K-41 and K-41Am were dissolved in dimethyl sulfoxide (DMSO) and then diluted with cell culture medium to the concentrations for the experiments. Our pervious data have shown that, when its concentration is 0.1% or lower, DMSO has little effect on the viability of the cells used in this study as well as HIV infection/replication in the cells (data not shown). In the final experiments, the highest concentration of DMSO was 0.025% and in the range of concentrations without cytotoxicity.
Viruses and cells
Two HIV-1 strains, HIV-1IIIB and HIV-1BaL (X4 and R5 virus, respectively), were used in this study [23]. Three type of cells (TZM-bl cells, MT-2 cells, and PBMCs) were used for HIV-1 infection/replication. HIV-1 replication was observed in the TZM-bl-HIV-1IIIB system, MT-2-HIV-1IIIB system, and PBMCs-HIV-1BaL system. HIV-1IIIB is a syncytium-inducing HIV-1 strain. MT-2 cell line, a human lymphocyte cell line, is suitable for X4 isolates [24]. Hela lineage TZM-bl cell line [25], expressing CD4, CXCR4, and CCR5, contains Tat-responsive reporter genes under the regulation of an HIV-1 long terminal repeat (LTR) [26]. PBMCs were from healthy blood donors and isolated from blood by centrifugation using Ficoll-Hypaque (GE Health Care, Uppsala, Sweden). TZM-bl cells were cultured in DMEM with 10% heat-inactivated foetal bovine serum (FBS), 100 IU/ml penicillin, 100 µg/ml streptomycin, and 1% L-glutamine. MT-2 cells and PBMCs were cultured in RPMI 1640 with 10% FBS, 100 IU/ml penicillin, 100 µg/ml streptomycin, 1% L-glutamine 50 ng/ml, and human interleukin-2 (IL-2, only for PBMCs). HIV-1IIIB, HIV-1BaL, MT-2, and TZM-bl cell lines were obtained from the Academy of Military Medical Sciences, China.
Cell viability assay
Cell viability was tested by the CellTiter-Glo® Luminescent Cell Viability Assay (Promega-Beijing, China) [27, 28]. The cells were mixed with serially diluted compounds in 96-well plates (Costar, Corning, USA) at a density of 1 × 104 cells/well (TZM-bl), 3.2 × 104 cells/well (MT-2), or 2 × 105 cells/well (PBMCs), respectively. After incubation in parallel with anti-HIV assays, cell viability assay reagent was added to cell cultures, and the luminescent signal was measured using a microplate reader (BioTek, Synergy H1, UK).
Anti-HIV assay
The anti-HIV assay was conducted in parallel with cell viability assay in mock-infected cell cultures. The anti-HIV assays were carried out using two HIV-1 strains and three types of cells described above; namely, three virus-cell systems (TZM-bl-HIV-1IIIB, MT-2-HIV-1IIIB, and PBMCs-HIV-1BaL system). To calculate IC50 and CC50 values, each compound was serially diluted to eight concentrations.
For TZM-bl-HIV-1IIIB system, TZM-bl cells (1 × 104 cells/well) were cultured for 24 h. The compounds and HIV-1IIIB virus (multiplicity of infection, MOI = 1) were added to the cultures. After 48 h, the virus levels were measured by the luciferase activity using the Promega Bright-Glo Luciferase Assay.
For the MT-2-HIV-1IIIB system, MT-2 cells (3.2 × 104 cells/well) were cultured for 24 h, the compounds and HIV-1IIIB virus (MOI = 1) were added to the cultures. After 72 h, 50 µl of culture supernatant from each well was transferred to TZM-bl cell cultures (2.7 × 104 cells/well) in 96-well plates. After 24 h, the levels of HIV-1 in TZM-bl cells were measured by the luciferase activity using the Promega Bright-Glo Luciferase Assay.
For PBMCs-HIV-1BaL system, prior to HIV-1 infection, the isolated PBMCs were stimulated with phytohemagglutinin (PHA)-P (5 µg/ml) for 72 h. Then the PBMCs (2 × 105 cells/well) were infected with HIV-1BaL (MOI = 0.01), and the compounds were added. After 5 days the culture supernatants were collected, and HIV-1 p24 levels in supernatants were determined using an HIV-1 p24 ELISA kit (Hebei Medical University Biomedical Engineering Centre, China).
The 50% inhibitory concentration (IC50) and 50% cytotoxic concentration (CC50) were calculated using non-linear regression and dose-response inhibition curves [29].
HIV-1 RT inhibition assay
The HIV-1 RT inhibition assay was performed using an ELISA RT assay kit (Roche, 11468120910, Germany) according to the manufacturer’s instruction [30]. The absorbance at 405 nm and 490 nm (A405/490) was determined using a microplate reader.
HIV-1 IN inhibition assay
The HIV-1 IN inhibition assay was performed using an HIV-1 Integrase Assay Kit (XpressBio, EZ1700, USA) according to the manufacturer’s instruction. Briefly, the streptavidin-coated plates were coated with a double-stranded HIV-1 LTR U5 donor substrate (DS) DNA, followed by the full-length recombinant HIV-1 integrase protein. Integrase testing reagents were then added, followed by a different double-stranded target substrate (TS) DNA containing a 3’-end modification. The products were colourimetrically measured using an HRP-labelled antibody against the TS 3’ end modification.
Ligand docking
Docking of K-41A and K-41Am to the targeted viral proteins was performed using the systemsDock (http://systemsdock.unit.oist.jp/) [31]. The procedure is described as follows:
Step 1: Specify the viral proteins and binding sites by PDB ID. The coordinates for HIV-1 reverse transcriptase (RT) core complex (PDB ID. 4R5P) [32] and HIV-1 integrase (IN) core complex (PDB ID: 1QS4) [33] were obtained from RCSB Protein Data Bank (http://www.rcsb.org/).
Step 2: The structures of two compounds were obtained from ChemBioOffice. Upload the structures of small molecules for the docking by uploading the structure files in the format of Mol2.
Step 3: Run docking simulation.
Step 4: Obtain simulation results.
Results
Anti-HIV-1 activities of K-41A and K-41Am
Dose-dependent inhibitory effects of K-41A and K-41Am on HIV-1 replication were all observed in the TZM-bl-HIV-1IIIB, MT-2-HIV-1IIIB, and PBMCs-HIV-1BaL systems (Figure 2). The anti-HIV activities of K-41A and K-41Am were rather strong. The IC50s of K-41A on HIV-1 replication in three systems were 0.75 µM, 0.09 µM, and 0.13 µM, respectively, and the IC50s of K-41Am were 5.57 µM, 0.24 µM, and 1.15 µM, respectively (Table I). Both compounds exhibited active anti-HIV status in different virus-cell systems, with selective indexes (SIs) > 5 (except K-41Am in TZM-bl-HIV-1IIIB) (Table I). The most potent activity was observed in K-41A in the PBMCs-HIV-1BaL system, with a highest SI of 598.00 (Table I).
Figure 2
Cytotoxicity and anti-HIV-1 activities of K-41A and K-41Am. The antiviral activities of K-41A and K-41Am were tested using three virus-cell systems, with cell viability measured in parallel mock-infected cells based on quantitation of the ATP activity. The cell viability of K-41A is shown in Figures 2 A–C, respectively. The cell viability of K-41Am is shown in Figures 2 G–I, respectively. In the TZM-bl-HIV-1IIIB system, the effects of K-41A (D) and K-41Am (J) on HIV-1 replication. In the MT-2-HIV-1IIIB system, the effects of K-41A (E) and K-41Am (K) on HIV-1 replication. In the PBMCs-HIV-1BaL system, the effects of K-41A (F) and K-41Am (L) on HIV-1 replication. The levels of HIV-1 BaL in the culture supernatants were determined by an HIV-1 p24 ELISA kit. The relative cell viabilities in compound-treated groups are expressed as the percentage of control (without compound treatment, which was defined as 100%). Relative virus replication levels are expressed as the percentage of control (with HIV-infection, without compound treatment, which was defined as 100%) The cell viability of K-41Am is shown in Figures 2 G–I, respectively. In the TZM-bl-HIV-1IIIB system, the effects of K-41A (D) and K-41Am (J) on HIV-1 replication. In the MT-2-HIV-1IIIB system, the effects of K-41A (E) and K-41Am (K) on HIV-1 replication. In the PBMCs-HIV-1BaL system, the effects of K-41A (F) and K-41Am (L) on HIV-1 replication. The levels of HIV-1 BaL in the culture supernatants were determined by an HIV-1 p24 ELISA kit. The relative cell viabilities in compound-treated groups are expressed as the percentage of control (without compound treatment, which was defined as 100%). Relative virus replication levels are expressed as the percentage of control (with HIV-infection, without compound treatment, which was defined as 100%)
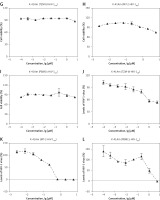
Table I
Summary of cell viability and anti-HIV-1 activities of K-41A and K-41Am
| Cells | Strains | Compounds | IC50a [µM] | CC50b [µM] | SIc |
|---|---|---|---|---|---|
| TZM-bl | HIV-1IIIB | K-41A | 0.75 | 4.36 | 5.81 |
| K-41Am | 5.57 | > 5.79 | > 1.04 | ||
| MT-2 | HIV-1IIIB | K-41A | 0.09 | 4.45 | 49.44 |
| K-41Am | 0.24 | 19.37 | 80.71 | ||
| PBMCs | HIV-1BaL | K-41A | 0.13 | 77.74 | 598.00 |
| K-41Am | 1.15 | > 5.79 | > 5.03 |
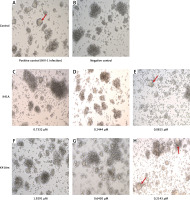
The protective effects of K-41A and K-41Am on HIV-1-induced cytopathic effect (CPE) were also observed (Figure 3). In the HIV-1 infection group (positive group), many syncytia formed (indicated by the arrows), representing the occurrence of virus-mediated cell-cell fusion (Figure 3 A). Whereas in negative controls, no syncytium was observed (Figure 3 B). When K-41A or K-41Am was presented at low concentrations, fewer syncytia were observed compared with those in positive control (Figures 3 E, H). When K-41A or K-41Am was present at higher concentrations, no syncytium was observed (Figures 3 C, D, F, G). These results indicate the dose-dependent inhibitory effects of K-41A or K-41 on the formation of syncytia induced by HIV-1 infection.
Figure 3
Protective effects of K-41A and K-41Am on HIV-1-induced cytopathic effect (CPE). MT-2 cells were infected with HIV-1IIIB. The infected cells were then treated with or without K-41A and K-41Am for 3 days. Microscopic observation (100×) of syncytia (indicated by the arrows) caused by HIV-1 infection. A – Positive control (HIV-1 infection, without compound treatment). B – Negative control (without HIV-1 infection and compound treatment). C–E – The HIV-1 infected cells were treated with K-41A at the shown concentrations. F–H – The HIV-1 infected cells were treated with K-41Am at the shown concentrations
K-41A and K-41Am inhibited the activities of HIV-1 RT and IN
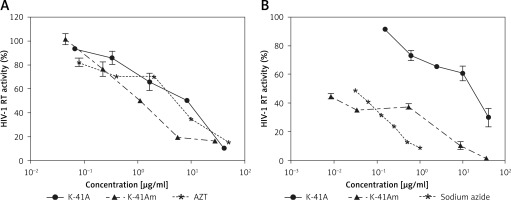
To illuminate the mechanism(s) involved in the anti-HIV-1 activity of K-41A or K-41Am, we focused on their effect on key HIV-1 enzymes and found that two compounds inhibited the activities of HIV-1 IN and RT (Figure 4 A). K-41A,K-41Am, and AZT (positive control) inhibited 50% of the RT activity at the concentration (IC50) of 4.48 µg/ml, 1.14 µg/ml, and 3.25 µg/ml, respectively (Figure 4 A). In addition, K-41A, K-41Am, and Sodium Azide inhibited 50% of the IN activity (IC50) at the concentration of 11.14 µg/ml and 0.01 µg/ml, and 0.03%, respectively (Figure 4 B). These results demonstrate that K-41A and K-41Am inhibited the activities of HIV-1 RT and IN.
Figure 4
K-41A and K-41Am inhibited the activities of HIV-1 RT and IN. The effects of K-41A and K-41Am on the activity of recombinant HIV-1 RT (A). The effects of K-41A and K-41Am on the activity of recombinant HIV-1 IN (B). HIV-1 RT and IN activity were measured by ELISA kits according to the protocol provided by the manufacturer. The HIV-1 RT activity or IN activity of control (without compound treatment) was defined as 100%. For HIV-1 RT activity assay (A), azidothymidine (AZT) treatment was used as a positive control. For HIV-1 IN activity assay (B), sodium azide treatment was used as a positive control
Docking models of HIV-1 IN and HIV-1 RT with K-41A and K-41Am
Molecular docking is widely used in quantitative structure-activity relationship (QSAR) studies, especially for the evaluation of anti-viral activity of newly discovered or synthesised compounds [30, 34]. Docking scores (pKd/pKi) usually ranging from 0 to 10 (i.e. from weak to strong binding) that represent the predicted binding affinity of K-41A or K-41Am to two targeted viral proteins are shown in an interactive table (Table II), a heat map (Figure 5 A), and a histogram (Figure 5 B). The protein residues involved in crucial contact with the compounds are also shown in Table II. As shown in Table II, compounds K-41A and K-41Am have similar docking scores on the viral proteins, with their enzyme residues more than native ligand. In addition, HIV-1 IN shows relatively high docking scores compared with HIV-1 RT. Therefore, these two compounds were considered to be potential active compounds targeting to two viral enzymes.
Figure 5
Docking simulation in SystemsDock. Docking scores (pKd/pKi) that represent the predicted binding affinity of K-41A and K-41Am to HIV-1 RT and IN. A – Heat map of the docking simulation. B – Histogram map of the docking simulation
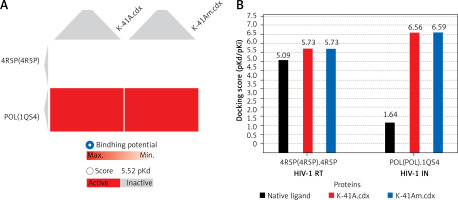
Table II
Docking scores (pKd/pKi) and residues of the docking simulation for each target protein and compound
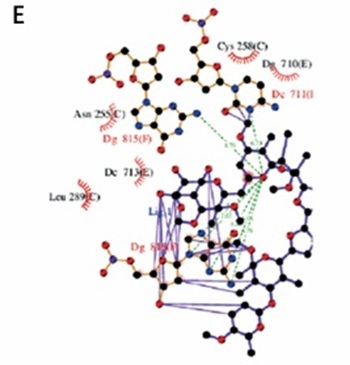
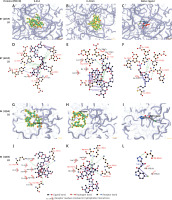
To better understand the interactions of two compounds with HIV-1 RT and IN, and to find some insights for structure optimisation, we analysed the binding modes. Figure 6 shows 3D and 2D representations of the lowest energy pose of K-41A, K-41Am and native ligand docked into the RT and IN binding pockets. Protein residues involved in the binding interaction are listed in 2D representations. For a reference, the binding interactions of the native ligand with proteins were also analysed and displayed. As shown in Figure 6, the ligand was accommodated through polar and hydrophobic interactions with pocket-lining residues. K-41A and K-41Am formed more hydrogen bonds with the target proteins than the native ligand formed. These results provide further evidence that K-41A and K-41Am could be viewed as promising compounds against HIV-1 RT and IN.
Figure 6
The detailed protein-ligand interactions for K-41A and K-41Am to HIV-1 RT and IN. 3D and 2D representations of the lowest energy pose of K-41A, K-41Am, and native ligand docked with the HIV-1 RT (PDB ID. 4R5P) and IN (PDB ID: 1QS4) binding pockets. A–C – 3D images of protein-ligand interactions of K-41A (A), K-41Am (B), and native ligand (C) with HIV-1 RT. D–F – 2D images of protein-ligand interactions of K-41A (D), K-41Am (E), and native ligand (F) with HIV-1 RT. G–I – 3D images of protein-ligand interactions of K-41A (G), K-41Am (H), and native ligand (I) with HIV-1 IN. J–L – 2D images of protein-ligand interactions of K-41A (J), K-41Am (K), and native ligand (L) with HIV-1 IN. In 3D images, the ligands are depicted as blue carbon sticks. HIV-1 RT and IN are shown as a transparent cartoon. The protein residues involved in crucial contacts with the compounds are indicated as carbon lines. In 2D images, protein residues involved in the binding interactions are indicated
Discussion
Screening of anti-HIV-1 compounds from natural marine actinomycete products has attracted increasing attention due to their unique chemical structures, rather than synthetic molecules [35–38]. In the present study, we used three virus-cell systems (TZM-bl-HIV-1IIIB, MT-2-HIV-1IIIB, and PBMCs-HIV-1BaL system) to characterise the inhibitory effects of two polyethers isolated from the marine-derived Streptomyces sp. SCSIO 01680, K-41A, and K-41Am on HIV replication in vitro. These two monoglycoside polyethers exhibited relatively strong anti-HIV-1 activity, with IC50 ranging from 0.09 µM to 5.57 µM (Table I). Mechanism research revealed that two compounds target two key viral enzymes, HIV-1 RT and IN, indicating that K-41A and K-41Am are actually multi-target compounds against HIV-1, which is in line with the current developmental trend of anti-HIV drug discovery. It is believed that multi-target anti-HIV drugs have a better therapeutic effect and less drug resistance than single-target drugs [15].
Currently, most FDA-approved anti-HIV-1 drugs inhibit HIV replication through blocking HIV-1 RT or IN [39]. HIV RNA is transcribed into its complementary DNA strand by HIV RT, followed by the viral DNA which is integrated into the host cell genome by HIV IN [40]. Therefore, targeting both HIV-1 RT and IN by the two polyethers may be the basis of their strong inhibitory activity against HIV-1 infection. In addition, the docking models of HIV-1 RT or IN with K-41A or K-41Am indicate that there were interactions between the compounds and two viral proteins, providing bioinformatics and structural evidence for the inhibitory activity of the two compounds.
Nevertheless, our finding is not the first to report that polyethers have anti-HIV-1 activity. An earlier study by Mariko Nakamura et al. indicated the inhibitory effects of 10 polyether antibiotics on HIV-1 replication [41]. They also reported relatively strong anti-HIV-1 activities of these polyether antibiotics, especially in chronical U937 cells, with IC50s ranging from 0.28 µg/ml to 10.1 µg/ml [41]. Their mechanism research shows that these polyethers suppressed HIV replication through affecting multiple phases of virus replication and infection cycles, and our results indicate that K-41A and K-41Am inhibit HIV-1 replication via targeting HIV-1 IN and RT. The difference in anti-HIV-1 mechanism between the earlier study and ours might be due to the different structures of polyethers. K-41A and K-41Am have a terminal (F) ring [22], which is rare in polyethers and was only previously found in K-41B [22] and CP-96,797 [21]. Thus, this rare terminal (F) ring may play an important role in the anti-HIV-1 activity of K-41A and K-41Am. In addition, our data show that K-41A was significantly stronger than K-41Am in inhibiting HIV-1 (Table I). The structure of K-41Am is very similar to that of K-41A. The only difference between the two compounds is the R group, K-41A is the H atom, and K-41Am is the methyl group, which means that this methyl group has a significant influence on the anti-HIV-1 activity of the compounds. This R group as well as the nearby structure will become important sites for structural modification to improve the anti-HIV activity.
In conclusion, our study demonstrates that two Streptomyces sp. SCSIO 01680-derived polyethers, K-41A and K-41Am, exhibited anti-HIV-1 activity. K-41A and K-41Am inhibit HIV replication through a multi-target inhibition mechanism, including suppressing the activities of HIV-1 IN and RT. These findings may provide a novel molecular pattern for anti-HIV drug design.