Introduction
There exist multiple risk factors in osteoporosis, for example age, bone mineral density (BMD), weight, bone geometry, caffeine [1, 2]. Osteoporosis may occur when the rates of formation and apoptosis become imbalanced in the highly specialized osteoblasts and osteoclasts involved in bone maintenance [3]. Although there are many studies on the pathways involving osteoblast differentiation and bone formation, the interactions between osteogenesis-related signaling pathways remain unclear. Therefore, it is vital to find novel regulatory mechanisms upstream of the signaling pathways. Metastasis associated lung adenocarcinoma transcript 1 (MALAT1), located on chromosome 11q13.1, is a long noncoding RNA (lncRNA) with vital biological functions. It was found that RANKL inhibits the proliferation of osteoblasts by upregulating the expression of MALAT1 in vitro [4]. In addition, MALAT1 was found to regulate the expression of β-catenin, and the overexpression and down-regulation of MALAT1 can respectively induce and decrease the expression of β-catenin [5, 6]. However, the effect of lncRNA MALAT1 on bone metabolism function is poorly understood.
LncRNA refers to a large class of endogenous RNA molecules that do not encode proteins; noncoding RNAs include microRNA and long noncoding RNA, among others. LncRNAs can interact with a miRNA, regulate its downstream target genes and be adsorbed by a “sponge” to bind homologous miRNA molecules competitively with the target mRNA through a microRNA response element (MRE), resulting in a relative decrease in miRNA level and activity. This inhibits the silencing effect of a miRNA on its target mRNA; furthermore, the expression level of the target gene can be upregulated and the posttranscriptional level of the gene can be regulated by lncRNA in the “ceRNA” mechanism [7, 8]. Screening of microarray data revealed that the miRNA miR-320a was upregulated after knockdown of MALAT1 expression, while database and literature results showed that miR-320a can reciprocally interact with MALAT1 [9]. A literature investigation found that miR-320a was overexpressed in osteoporosis samples. miR-320a can target and inhibit CTNNB1, the gene of catenin β1, and subsequently inhibit lymphoid enhancer-binding factor 1 (LEF1) and RUNX2; therefore, it is considered that miR-320a has the effect of inhibiting osteogenesis differentiation [10]. The β-catenin gene was shown to be a direct target gene of miR-320a, and miR-320a negatively regulated the expression of β-catenin in cancer research [11]. In addition, studies have shown that miR-320a can target and inhibit the expression of NRP-1 [12].
A study found that inhibition of NRP-1 could affect the differentiation of osteoblasts by inhibiting the Wnt/β-catenin signaling pathway [13]. NRP-1, which is not only expressed on the surface of osteoclast precursors and osteoclasts but also expressed on the membrane of osteoblasts, plays a critical role in the molecular basis of Sema3A as a double regulator and modulator of bone [14, 15]. The NRP-1 and PlexinA1 receptor complex, which can implement cotransduction of intracellular signals after binding Sema3A, has the unique dual functions of promoting osteogenesis and inhibiting osteoclast differentiation, functions that are of great significance for promoting both bone formation and balancing bone metabolism [15]. Therefore, we speculate that MALAT1 may regulate the expression of NRP-1 and β-catenin by regulating miR-320a in the competing endogenous regulation mechanism.
The potential inherent involvement of MALAT1 in osteoporosis is investigated in the present investigation. In addition, we examined the mRNA expression level of miR-320a and MALAT1 in bone mesenchymal stem cells (BMSCs) derived from osteoporotic rats, and its effects on osteogenic differentiation and cell growth were tested. Furthermore, the underlying regulatory mechanism of the microRNA miR-320a in BMSCs were analyzed in vitro. Finally, the possible interaction between miR-320a and β-catenin, miR-320a and NRP-1 was also explored in BMSCs. The present investigation may promote better understanding of the regulatory mechanisms and correlations of MALAT1/miR-320a/NRP-1/β-catenin in osteogenesis in vitro.
Material and methods
Experimental animals
Female Sprague Dawley (SD) rats (n = 51), aged 3 months, weighing 226 ±22 g, were purchased from the Chengdu Laboratory Animal Center (Chengdu, China). From them, 48 female rats were divided into control (n = 24) and model (n = 24) groups at random, while 3 female rats were used for molecular experiments. Three animals were kept in each cage in a 25°C temperature-controlled room with 50–60% relative humidity and a 12 h light and 12 h dark cycle.
Establishment of the OVX animal models
After acclimatization for 2 weeks, following the intraperitoneal injection of ketamine (3B Scientific Corporation, USA) at a dose of 100 mg/kg, the rats underwent bilateral ovariectomy (OVX, n = 24) or sham surgery (Sham, n = 24). To obtain the tibial bone samples, the animals were sacrificed with euthanasia 3 months after the surgery.
BMSC isolation and primary cell culture
BMSCs were isolated from the femurs and tibiae of SD rats (n = 3), and we cultured the BMSCs according to previously reported methods [16]. Cells were incubated in 5% CO2 in 95% humidity at 37°C. The culture media consisted a rate of 10 : 1 : 0.1 minimum essential medium α medium (α-MEM, Gibco, USA), 10% fetal bovine serum (FBS, Gibco, USA), and penicillin-streptomycin (Gibco, USA). When adherent cells reached 90% confluence, the cells were treated with a 0.25% trypsin solution for continued passage. BMSCs, derived from the third generation, were prepared for the subsequent experiments. The transfected BMSCs for each group were grown for approximately 7 days, and the culture wells were supplemented with osteogenesis inducing conditional medium. Osteogenic induction medium contains 500 ml of α-MEM, 50 g/ml ascorbic acid, 10–8 M dexamethasone, and 8 mM β-glutamine.
Hematoxylin and eosin (HE) staining and micro-computed tomography (micro-CT) scanning
The tibial metaphyseal bone of ovariectomy and sham-operated rats was fixed with 10% polyformaldehyde. The tibial metaphyseal bone underwent 5% nitric acid decalcification, washing with flowing water, routine dehydration, and xylene permeabilization. Tibial axis sections (5 μm) were obtained, hematoxylin and eosin staining was performed, a neutral resin seal sheet was applied, tissue sections were observed under an optical microscope to analyze the morphology of the bone and tissue, and microarchitecture of the proximal rat tibiae was examined by micro-computed tomography (micro-CT) scanning by a CT scanner (Scanco Medical, Bassersdorf, Switzerland) to determine whether the model of osteoporosis was successful. The resolution for the slices was 2048 × 2048 pixels, and the isotropic voxel size was 10 μm. A CT system (70 kV, 114 μA; μ-CT 80 scanner) was used to scan the volume of interest. To evaluate the bone morphometric parameters to the proximal tibial metaphysis, we defined the volume of interest (VOI) as the distal slices (n = 100) from the proximal tibial growth plate.
Quantitative reverse transcription polymerase chain reaction (RT-PCR) assays
Quantitative RT-PCR was used to determine the level of miR-320a and MALAT1 in the bone tissue of each group; MALAT1 siRNA interference efficiency, transfection efficiency, and mRNA expression in Osterix and Runx2 were also evaluated.
To explore the expression levels of miR-320a and MALAT1 in the bone tissue of the osteoporosis group and the control group, tibial bones of the ovariectomy and control groups were harvested 3 months after the ovariectomy or sham surgery. The tibial metaphysis was placed in a TRIzol (1 ml) (Gaithersburg, MD) homogenate tube for shredding and grinding to extract the total RNAs; according to the instructions of the manufacturer, RNA was extracted step by step and used for amplification. The subsequent steps included denaturation and amplification.
After 14 days of osteogenic induction, the cultured and harvested BMSCs were flushed with TRIzol reagent (1 ml) (Gaithersburg, MD), and the RNA and reaction system was prepared according to the manufacturer’s specification. Melting curve analysis was performed after the process of amplification and the subsequent fluorescence measurements. For the quantification of all mRNA expression levels, control wells without cDNA were induced for analysis using the 2–ΔΔCt method to analyze the fold changes in miRNA or the related gene expression changes in each sample. The primer (Carlsbad, CA, USA) sequences, including the sequence for MALAT1, RUNX2, Osterix, miR-320a and U6, used for RT-PCR are listed in Table I (when GAPDH was used for normalization of data).
Table I
PCR primers used for PCR
Small interfering RNA
Three kinds of small interfering RNAs (siRNA) targeting MALAT1 and miR-320a mimics and the negative control were prepared by 100BIOTECH (Hangzhou, China). They were transfected to BMSCs with Lipofectamine 2000 reagent according to the manufacturer’s instructions. Table II lists the primer sequences of the siRNA and control siRNA for MALAT1. The process of transfection of siRNA into BMSCs is based on the manufacturer’s instructions. New medium was added to the cells after transfection for 24 h, and the in vitro interference efficiency of four groups of siRNA was determined by qRT-PCR.
Table II
Paired primers of siRNAs and NC
Immunohistochemical (IHC) analysis
The paraffin-embedded tissue samples (tibial bone from ovariectomy and sham surgery rats) were examined to determine the protein expression of NRP-1 and β-catenin. Specimens were blocked with 4% paraformaldehyde (24 h) and decalcified in 10% EDTA (Nanjing BaiRui Biotechnology Co. Ltd.) solution (1 month). After gradient ethanol dehydration and baking, paraffin sections were obtained. Sections were heated to a boil, then heated at low heat for 20 min for antigen retrieval. Sections were placed in 3% H2O2 solution and incubated at room temperature for 10 min to block endogenous peroxidase. Then, sections were washed 3 times with PBS (5 min each time), dried and blocked in 5% BSA for 20 min. After the removal of BSA liquid, 50 μl of primary antibody diluted against NRP-1 and β-catenin (rat polyclonal anti-NRP-1 antibody 1 : 250, rat polyclonal anti-β-catenin antibody 1 : 500, Boster, Wuhan, China) was added to each section, and the tissues was covered at 4°C overnight. The rat peroxidase-diaminobenzidine kit (Maixin_Bio, Fuzhou, China) was used in this experiment. To each slice 50–100 μl of biotinylated second antibody (goat anti-rat, dilution ratio 1 : 1000) of the corresponding species was added. Each section received 50–100 μl of freshly prepared DAB solution (Beyotime, Suzhou, China), stained with hematoxylin, rinsed and turned blue. The cells that stained brown were regarded as positive cells. Five high power microscopic fields were selected from each section from the inverted fluorescence microscope to count positive cells at random, and finally the average value was obtained.
Osteogenic differentiation induction
After transfection, the BMSCs in each group grew for approximately 7 days, the osteogenic induction medium (containing 500 ml of α-MEM, 50 g/ml ascorbic acid, 10–8 M dexamethasone, and 8 mM β-glutamine), obtained from Sigma (St. Louis, MO), was added to the culture wells. The alkaline phosphatase (ALP) activity of osteoblasts was determined by using an alkaline phosphatase kit after 14 days of induction.
CCK-8 kit assay
A CCK-8 Kit (Beyotime, Suzhou, China) was used to determine the changes in cell activity in each group on the 14th day after culture. Ten microliters of CCK-8 solution was added to each well for each group at 37°C and 2 h, the sample was incubated with 5% CO2, and the OD values were calculated at 450 nm with ELISA (BioTek, USA) on the 14th day of culture.
Alkaline phosphatase (ALP) activity
The ALP activity of osteoblasts was determined by an ALP kit 14 days after induction. The working solution of the standard product was prepared with 15 μl of p-nitrophenol phosphate (pNPP) solution (10 mM) and was diluted to 0.3 ml to reach a final concentration 0.5 mM. The ALP activity was calculated by using the OD value at 405 nm per milligram of total protein. All the experiments were conducted in triplicate.
Alizarin red staining
Each group of BMSCs grew for approximately 7 days, and culture wells were supplemented with the osteoblast-inducing medium. The culture medium was replaced every 2 days to observe the growth and calcification of the osteoblasts. After 21 days of continuous culture, alizarin red staining was performed to evaluate the calcium deposition/mineralization of osteoblasts. The culture supernatant was washed with PBS 3 times. Alizarin Red S dye solution (Wuhan, China) was added at 37°C. The microscope was used to obtain images and records of the plates.
Western blot assay
Each cell sample was divided into two parts and one part was used to determine the expression of β-catenin protein by using a cytoplasmic and nuclear isolation kit to extract cytoplasmic and nucleoproteins, respectively. Total protein was collected to determine the quantitative protein levels of neuropilin-1 (NRP-1), osteocalcin (OCN), and osteopontin (OPN). BCA quantification was performed strictly according to the instructions of a BCA protein concentration assay kit. The membrane was sealed with secondary antibody (10% goat serum). Primary antibody (1 : 1000, rat anti-NRP-1, OCN, OPN IgG, Beyotime, China) was added to the membrane and incubated for 1 night at 4°C. After being washed with PBS three times, the secondary antibody (1 : 4000, goat anti-rat antibody, Beyotime, China) was added to the membrane and incubated at 37°C for 1 h.
Statistical analysis
Data were analyzed by SPSS 16.0 (SPSS, Chicago, IL), and the results are expressed as the mean ± SD. In the animal study, specimens were harvested from 48 different rats for each group. For cell analysis, all experiments were repeated 3 times in triplicate. p < 0.05 indicated a statistically significant difference. Comparison between two groups was performed using the T-test, and comparison among four groups was conducted by one-way ANOVA.
Results
Evaluation of the bone specimens from OVX and sham-operated rats
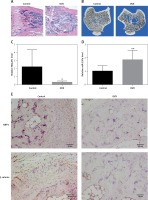
Tibia specimens were harvested from OVX and sham-operated groups 3 months after the operation. The pathological changes in the tibia examined by using histological sections and micro-CT scanning confirmed the establishment of osteoporosis (Figures 1 A, B). The trabeculae of the sham-operated group were uniform in size, arranged and connected into a net. In the OVX group, the trabeculae were broken, the structures were distorted and loosely arranged, and most of them could not be connected into a network (Figure 1 A). Quantitative micro-CT analysis showed an obvious decrease in percent bone volume, as well as damaged bone microarchitecture, in the OVX group compared to that in the control group (Table III). The trabecular separation for OVX rats increased 380% compared with the sham operated groups (p < 0.01, Table III). Compared with the sham operated groups, Tb·Th decreased by 46.1% (p < 0.05) and Tb·N decreased by 69.6% (p < 0.05, Table III). RT-PCR results showed that the MALAT1 expression in the bone tissue of the OVX group was decreased by 85.6% (p < 0.05) (Figure 1 C), while the expression of miR-320a was increased by 82.9% (p < 0.01) compared to that of the control (Figure 1 D). The IHC staining showed more positive brown color in the femur of rats in each sham operated group, and the IHC results showed inhibited expression of NRP-1 and β-catenin in OVX rats compared to that in the control (Figure 1 E). The number of NRP1 positive cells expressed in the osteoporotic group compared with the control group was significantly decreased (p < 0.05, Table IV). Compared with the control group, the positive number of β-catenin expressed in the osteoporotic group was significantly decreased (p < 0.05, Table IV).
Table III
Parameters used in micro-CT to evaluate microstructure changes in tibia
| Microstructure parameter | Sham-operated | OVX |
|---|---|---|
| BV/TV | 0.43 ±0.12 | 0.13 ±0.08** |
| Tb·Sp [mm] | 0.21 ±0.04 | 1.01 ±0.09** |
| Tb·Th [1/mm] | 0.13 ±0.03 | 0.07 ±0.01** |
| Tb·N [1/mm] | 3.45 ±0.41 | 1.05 ±0.22** |
Table IV
Number of NRP1-positive and β-catenin-positive cells after immunohistochemistry in the tibia (1000×)
| Group | N | Control | Model | P-value |
|---|---|---|---|---|
| NRP1 | 24 | 31.3 ±5.2 | 13.6 ±3.1 | 0.021 |
| β-catenin | 24 | 29.7 ±4.7 | 10.1 ±1.4 | 0.017 |
Figure 1
A – HE staining results of the tibial metaphyseal bone obtained from ovariectomy and sham surgery rats (n = 24). B – Images from micro-CT scanning of the proximal tibia in osteoporotic and intact SD rats; a significant reduction in the number of trabeculae and a decrease in the continuity of bone trabeculae can be seen in the osteoporosis group. C, D – Expression levels of miR-320a and MALAT1 were determined by qRT-PCR, and significantly reduced expression of MALAT1 and overexpression can be seen in Figure 1 C and 1 D. E – Protein expression of NRP-1 and β-catenin was determined by immunohistochemistry. Scale bar = 20 µm (*p < 0.05, **p < 0.01 compared with control)
Effects of MALAT1 interference on miR-320a expression and cell activity in BMSCs
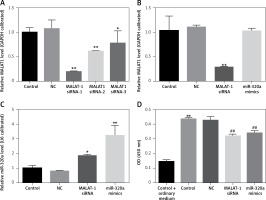
After transfection for 48 h, the transfection efficiency of MALAT1 siRNA and miR-320a mimics was confirmed by RT-PCR evaluation (Figures 2 A, B). According to Figure 2 A, we found that MALAT1 siRNA-1 had the best interference efficiency, so we selected MALAT1 siRNA-1 for our further experiments (Figure 2 A).Compared to the control group, the group with MALAT1 interference had increased expression of miR-320a (Figure 2 B), while overexpression of miR-320a showed no effect on the MALAT1 level (Figure 2 A). The CCK8 test was conducted after transfection for 14 days. Compared to cells cultured in ordinary medium, the cells in osteogenic medium had increased cell activity; for osteogenesis induced cells, gene transfection of MALAT1 siRNA and miR-320a mimics decreased the cell activity, as shown by the CCK-8 kit results 14 days after osteogenic induction.
Figure 2
A – MALAT1 level in BMSCs was determined by qRT-PCR after transfection with the vector, MALAT1 siRNA-1, MALAT1 siRNA-2 and MALAT1 siRNA-3 for 48 h. MALAT1 siRNA-1 showed the best transfection efficiency and can be used for further study. The MALAT1 level (B) and the miR-320a level (C) in BMSCs were determined by qRT-PCR after transfection with the vector, MALAT1 siRNA-1, and miR-320a mimics for 48 h. D – Activity of BMSCs was observed via a CCK-8 assay after osteogenesis was induced with osteogenic induction medium for 14 days. The groups in the CCK-8 test were divided into the following groups: 1) Control + ordinary medium: blank control + culture media; 2) Control: blank control + osteogenic induction medium; 3) NC: vectors + osteogenic induction medium; 4) MALAT1 siRNA: MALAt1 siRNA1 transfection + osteogenic induction medium; 5) miR-320a mimics: miR-320a mimics transfection + osteogenic induction medium (*p < 0.05, **p < 0.01 compared with control)
ALP activity, RT-PCR, and calcified nodule evaluation of osteogenic differentiation of BMSCs
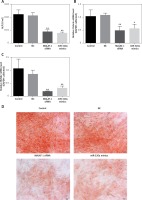
Fourteen days after inducing culture, the ALP activity of induced BMSCs in the MALAT1 siRNA group and the miR-320a mimic group was decreased by 60.9% and 66.4% respectively, compared to that in the control (p < 0.01); the above results showed that Malat1 siRNA group and miR-320a mimics had weaker bone formation ability compared with control group (Figure 3 A). The expression of Osterix was decreased by 53.2% (p < 0.01) in the MALAT1 group and by 46% (p < 0.05) in the miR-320a mimics group (Figure 3 B). The expression of Runx2 was decreased by 83% in the MALAT1 siRNA group (p < 0.01) and by 70.5% in the miR-320a mimics group (p < 0.01) compared to that in the control group (Figure 3 C). As a measure of critical functions in the mature osteoblasts, the matrix mineralization and calcium deposition levels were evaluated by Alizarin red staining under an inverted microscope after 21 days of osteogenetic induction. The number of orange calcified nodules observed in the MALAT1 siRNA group and miR-320a mimics group decreased significantly compared with that in the control group (Figure 3 D).
Figure 3
A – ALP activity of osteoblasts was determined by alkaline phosphatase kit 14 days after osteogenesis induction in BMSCs (ALP: alkaline phosphatase), and the activity of ALP in the MALAT1 siRNA group and miR-320a mimics group was significantly decreased (p < 0.01). The expression of Osterix (B) and RUNX2 (C) was determined by qRT-PCR after osteogenesis was induced in BMSCs for 14 days (RUNX2: Runt-related transcription factor 2). D – Alizarin red staining was conducted to observe the formation of calcified nodules under an inverted microscope after BMSC osteogenesis was induced for 21 days; a large number of calcified nodules were observed in the control group and NC group compared with the MALAT1siRNA group and miR-320a mimics group, and the number of calcified nodules in the cells of the MALAT1 siRNA group and miR-320a mimics group was significantly decreased. Scale bar = 20 µm (*p < 0.05, **p < 0.01 compared with control)
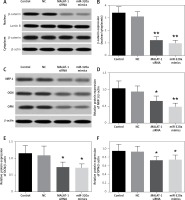
Western blot assay of β-catenin, NRP-1, OCN and OPN in BMSCs
The cytoplasm and nuclear fraction of β-catenin protein were extracted by cytoplasm and nuclear isolation kits, respectively, and then evaluated by a Western blot assay (Figure 4 A). The β-catenin level in the cytoplasm showed no difference, but that in the nucleus decreased significantly in the MALAT1 siRNA and miR-320a mimics groups. The ratio of the nuclear to cytoplasmic β-catenin in the MALAT1 siRNA group and the miR-320a mimics group showed significant downregulation by 65.1% (p < 0.01) and 72.2% (p < 0.01) compared to that in the control (Figure 4 B). NRP-1, OCN and OPN were also measured by Western blotting after 21 days of osteogenic induction (Figure 4 C). The results demonstrated that the expression of NRP-1, OCN and OPN protein was significantly decreased in the MALAT1 siRNA group compared to that in the control by 36.8% (p < 0.01), 35.6% (p < 0.05) and 23.1% (p < 0.05) respectively. Similarly, the expression of NRP-1, OCN and OPN protein in the miR-320a mimics group also decreased by 53.4% (p < 0.01), 42.9% (p < 0.05) and 21.2% (p < 0.05), respectively, compared to that in the control group (Figures 4 D–F).
Figure 4
A – Protein level of β-catenin in the cytoplasm and nucleus was determined by Western blot; β-catenin expression was observed to be significantly decreased in MALAT1 siRNA and miR-320a mimics groups in the nucleus. B – The ratio of β-catenin in the nucleus to that in the cytoplasm was calculated and compared between groups (**p < 0.01 compared with control). C–F – Western blot analysis was used to identify the osteogenic differentiation of BMSCs in different groups, by evaluation of the protein levels of NRP-1, OCN, and OPN (NRP-1 – neuropilin-1; OCN – osteocalcin; OPN – osteopontin) (*p < 0.05, **p < 0.01 compared with control)
Discussion
There may be a correlation between osteoporosis and other diseases such as left ventricular hypertrophy (LVH) [17]. Osteoporosis mainly results from an imbalance between osteoblastogenesis and osteoclastogenesis. The process in which osteoblasts are derived from multipotent MSCs plays a critical role in the development of osteoporosis and involves a complex of molecules that mediate cell-matrix and cell-cell interaction [3]. Over time, studies focused on the underlying regulatory mechanism of osteoporosis have shifted from protein-coding RNA to noncoding RNAs.
LncRNAs, which mediate epigenetic modifications of DNAs but do not encode proteins, are a group of transcripts that are more than 200 nucleotides long [18]. The evidence for lncRNAs in osteoporosis is accumulating, and lncRNA are believed to be involved in the osteoblastic differentiation [19]. Moreover, 59 differentially expressed lncRNAs were found to be upregulated and 57 were downregulated between untreated and BMP2-treated groups of MSCs undergoing osteoblast differentiation [20]. However, the underlying role that lncRNA plays in osteoporosis remains to be further delineated. In previous studies, the lncRNA MALAT1 was found to be upregulated in osteoblast cells in vitro; the lncRNA MALAT1 was closely related to the process of RANKL binding to the receptor RANK, and MALAT1 expression can be induced by RANKL [4].
An experiment showed that local use of medicine at the site of post-menopausal osteoporosis in vivo helped to promote the osseointegration [21]. It has been reported that MALAT1 can directly interact with miR-204 and functions as a miRNA sponge to positively regulate the osteogenic differentiation involved in CAVD (calcific aortic valve disease) pathogenesis [22]. Nevertheless, little is clear about the function of MALAT1 in osteoporosis and the underlying role of MALAT1 in bone metabolism. In the present study, we focus on MALAT1, which was investigated in cancer and tumor and is known as a noncoding RNA that is spliced infrequently. Through a rat model of osteoporosis, we observed that the level of MALAT1 in bone tissue was dramatically decreased in osteoporotic rats compared with that in control rats. We found that the osteoblasts induced from transfected BMSCs treated with MALAT1-siRNA showed significant inhibition of cell viability, calcium nodule formation, alkaline phosphatase activity and deposition of calcium nodules. Therefore, we further speculated that MALAT1 has an effect on osteogenic differentiation in bone metabolism and that the increase of MALAT1 in BMSCs may be related to the development of osteoporosis.
miRNAs are approximately 22 nucleotides long and can exert a silencing effect by mediating posttranscriptional modification to regulate the process of generating protein from mRNA [23]. MicroRNAs have critical roles in different diseases, they can mediate the metabolism of statins through the way of inflammation modulation and immune regulation in cardiovascular disease (CVD) [24]. MicroRNAs are new promising therapeutic targets for heart failure and familial hypercholesterolemia (FH) patients [25, 26]. It has been emphasized that abnormalities in the network of microRNA signaling may promote both the development and progression of osteoporosis, and, more importantly, miRNAs are emerging as crucial modulators of bone mass maintenance in osteoporosis [27]. However, further work to validate the underlying mechanisms before making definitive conclusions is required. A recent review highlighted both the advantages and challenges of using miRNAs in the circulating system as potential biomarkers, such as in patients with bone disease [28]. Competitive endogenous RNA (ceRNA) has specific mechanisms for regulating the influence of long noncoding RNAs (lncRNAs) on mRNAs levels by competing for a limited pool of microRNAs [29]. miRNAs act mainly to control gene downregulation by silencing the mRNA translation process after a specific sequence binds to its target gene through microRNA recognition elements (MRE), and miRNAs may also bind lncRNA species [8, 29]. MALAT1 can act as a miRNA sponge to regulate the expression of target miRNAs (including miR-320a) and protein-coding genes as a hypothetical type of competitive endogenous RNA (ceRNA) [9]. A recent study proved that upregulation of the microRNA miR-320a can negatively regulate the levels of MALAT1 and that miR-320a can reciprocally interact with MALAT1 [9]. However, it remains to be elucidated whether MALAT1 can regulate miR-320a to promote osteoblast differentiation from BMSCs through the ceRNA mechanism in vitro.
Dramatically increased MALAT1 expression was observed in osteoporotic rats compared to that in control rats in bone tissue examined by qRT-PCR. According to the present research, miR-320a was identified as a suppressor that inhibited the proliferation and osteogenic differentiation of BMSCs. Here, it was found that the level of miR-320a was significantly upregulated in BMSCs with MALAT1 silencing, but no obvious expression change in MALAT1 was found in the miR-320a mimics group; that is, the downregulation of MALAT1 can upregulate the level of miR-320a in BMSCs. However, the increase in miR-320a had no significant effect on the MALAT1 expression in BMSCs. Furthermore, the same phenomenon was observed in the BMSCs of the miR-320a mimics group. The increased miR-320a level in BMSCs also inhibited the activity of osteoblasts, the deposition of calcium nodules and the activity of alkaline phosphatase, which we had observed in the MALAT1 siRNA group in vitro. In other words, we proved that MALAT1 repression regulates the osteogenic differentiation of BMSCs through an effect mediated by miR-320a, which is mediated by the ceRNA mechanism. However, whether proteins and osteoblast-related pathways are involved in the differentiation process that is regulated by ceRNA remained unclear. Therefore, we further investigated some key proteins and pathways related to osteogenesis.
Previous studies have shown that OPG promotes bone formation through inhibiting bone resorption and RANKL inhibition in an OVX rat models [30]. Runx2 is upregulated first, followed by Osterix, and ALP, OCN, and OPN are then upregulated as late protein markers [31]. We found that decreasing MALAT1 or increasing miR-320a can inhibit the osteogenic differentiation of rat BMSCs by inhibiting the mRNA level of Runx2 and Osterix, as determined by RT-PCR. OPN can be produced by cells such as preosteoblasts and osteoblasts and plays major roles in bone morphogenesis, immunoregulation and inflammation [32]. One study reported that the increase in OCN was associated with an increased level of new bone formation and osteocalcin, an early marker of osteogenesis, in rat mesenchymal stem cells [31, 32]. Our in vitro evaluation of OCN and OPN verified that the decrease in osteogenic activity of BMSCs could be due to an increased level of miR-320a resulting from the decrease in the MALAT1 level.
NRP-1 and Sema3A binding can stimulate the differentiation of osteoblasts or inhibit the differentiation of osteoclasts respectively, through the canonical Wnt/β-catenin signaling pathway, and these effects are mediated by the receptor activator of nuclear factor-κB ligand (RANKL) secreted by osteoblasts [15]. It has been found that miR-320 can negatively regulate the expression of NRP-1 through regulating the target gene of NRP-1, contributing to the proliferation and migration of CCA cells [12]. We confirmed that the knockdown of MALAT1 significantly inhibited the expression of NRP-1 protein through the ceRNA mechanism in vitro. Canonical Wnt/β-catenin signaling plays a critical role in the regulation of bone homeostasis, while activation of this signaling can lead to increased bone mass and strength and is an intervention target for restoring mineral density [33]. The β-catenin signaling pathway positively regulates osteoblasts by activating T cell factor (TCF)-dependent transcription [34]. Studies have found that upregulated MALAT1 can modulate Wnt/β-catenin signaling to inhibit oral tongue squamous cell apoptosis. Furthermore, evidence has shown that downregulation of the lncRNA MALAT1 can decrease β-catenin expression in esophageal cancer cells in vitro [6]. β-Catenin can target miR-320a directly, and β-catenin overexpression is associated with low expression of miR-320a in glioblastoma multiform (GBM) cells [35]. In addition, it has been proved that β-catenin can target and regulate miR-320a directly in colon cancer cells [27].
The results of the experiment show a decrease of MALAT1 in osteoporotic rats, which is consistent with the decrease of osteogenic differentiation in rats after interfering with MALAT1. Therefore, we speculate that the intervention of MALAT1 can regulate the osteogenic differentiation of BMSCs in osteoporotic patients. This experiment is helpful to understand the potential regulatory mechanism of osteoporosis patients. MALAT1 may be a potential target for the treatment of osteoporosis.
In the present research, we found that a decrease of MALAT1 or an increase in miR-320a significantly reduced the nuclear β-catenin in osteoblasts; however, there was no reduction in the cytoplasm. It can be speculated that the distinct difference in inhibitory effects between the cytoplasm and nucleus may be related to the location of MALAT1, which is a common phenomenon in the “competitive endogenous RNA” hypothesis, where a preferential competitive combination exists in neighboring connections. In general, we speculate that due to the low expression of MALAT1, miR-320a upregulation followed the downregulation of β-catenin and NRP-1 expression through the mechanism of ceRNA, leading to the osteogenic differentiation inhibition of BMSCs in osteoporosis patients. More importantly, our study may open the road to new gene regulatory networks acting through miRNA-lncRNA competitive interactions in osteoporosis. We first demonstrated that NRP-1 and β-catenin can be functionally targeted by miR-320a directly in BMSCs and highlighted the potential value of lncRNA MALAT1 and miR-320a as promising targets for a novel treatment of osteoporosis.
In conclusion, this knowledge of MALAT1/miR320a in BMSC function may shed light on mechanisms underlying osteoporosis. The upregulated miR-320a expression may result from the down-regulated MALAT1 expression, which can be explained by the ceRNA mechanism.