Current issue
Archive
Manuscripts accepted
About the Journal
Editorial office
Editorial board
Section Editors
Abstracting and indexing
Subscription
Contact
Ethical standards and procedures
Most read articles
Instructions for authors
Article Processing Charge (APC)
Regulations of paying article processing charge (APC)
NEPHROLOGY / CLINICAL RESEARCH
Metabolic profiles of IgA nephropathy, membranous nephropathy, and diabetic nephropathy
1
Department of Nephrology, Fifth Hospital of Shanxi Medical University (Shanxi Provincial People’s Hospital), Taiyuan, China
2
Department of Medicine, International College of Medicine and Pharmacy, Changsha Medical University, Changsha, China
3
Key Laboratory of Kidney Disease, Precision Medicine Center, Shanxi Provincial People’s Hospital, Shanxi Medical University, Taiyuan, China
Submission date: 2025-02-12
Final revision date: 2025-08-10
Acceptance date: 2025-09-12
Online publication date: 2026-02-08
Corresponding author
Rongshan Li
Department of Nephrology Fifth Hospital of Shanxi Medical University (Shanxi Provincial People’s Hospital) Key Laboratory of Kidney Disease Precision Medicine Center Shanxi Provincial People’s Hospital Shanxi Medical University Taiyuan, China
Department of Nephrology Fifth Hospital of Shanxi Medical University (Shanxi Provincial People’s Hospital) Key Laboratory of Kidney Disease Precision Medicine Center Shanxi Provincial People’s Hospital Shanxi Medical University Taiyuan, China
KEYWORDS
TOPICS
ABSTRACT
Introduction:
Immunoglobulin A nephropathy (IgAN), membranous nephropathy (MN), and diabetic nephropathy (DN) are prominent contributors to chronic kidney disease burden. Our main objective was to contribute to the understanding of the metabolic profiles of these three major types of nephropathies and identify potential metabolic biomarkers.
Material and methods:
Kidney samples of 20 sex- and age-matched patients with biopsy-proven IgAN, MN, DN, and controls without any kidney diseases were included. Ultra-high-performance liquid chromatography–mass spectrometry analysis was conducted. The t-test was used to evaluate the statistical significance of the identified metabolites. Metabolic pathways were analyzed using the Kyoto Encyclopedia of Genes and Genomes (KEGG). Specificity, sensitivity, and area under the curve (AUC) were calculated to evaluate the predictive performance of metabolites.
Results:
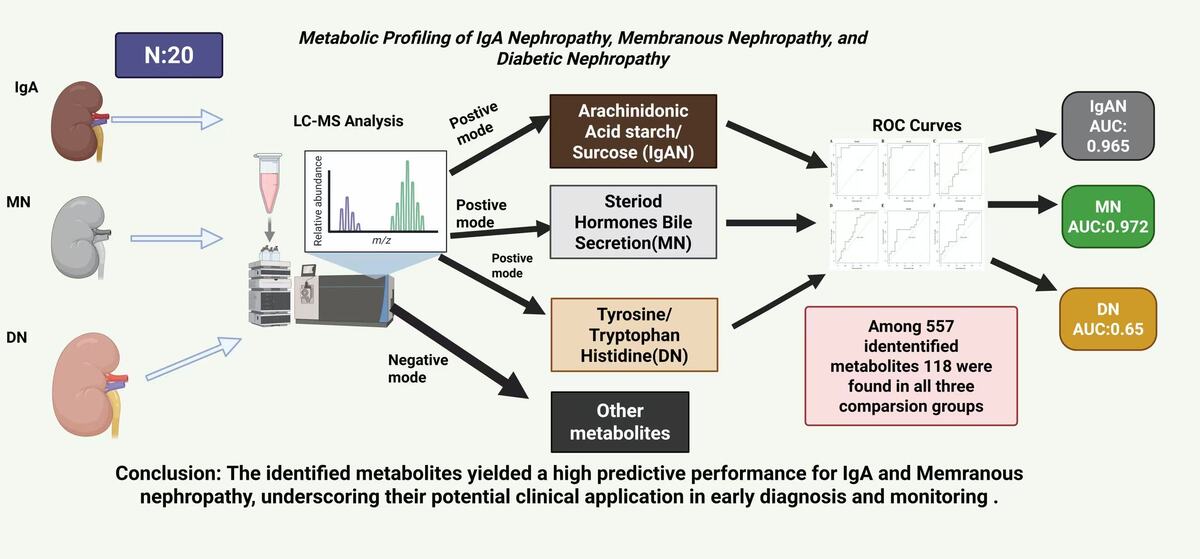
Among 557 identified differential metabolites, only 118 were found in all three comparison groups. Differential metabolites of IgAN vs. controls were significantly enriched in arachidonic acid metabolism, starch and sucrose metabolism, ferroptosis, and other pathways. In the DN group, metabolites were mainly enriched in phenylalanine, tyrosine and tryptophan biosynthesis, histidine metabolism, etc. MN-enriched pathways included steroid hormone biosynthesis, neuroactive ligand-receptor interaction, and bile secretion. In the positive mode, cumulative AUC values for comparison pairs IgAN vs. controls, MN vs controls, and DN vs controls were 0.965, 0.972, and 0.573, respectively, whereas in the negative mode the AUC values of all three pairs were slightly above 0.65.
Conclusions:
IgAN, MN, and DN have similar but distinct metabolic profiles. Only positive node metabolites of IgAN and MN exhibited high predictive performance.
Immunoglobulin A nephropathy (IgAN), membranous nephropathy (MN), and diabetic nephropathy (DN) are prominent contributors to chronic kidney disease burden. Our main objective was to contribute to the understanding of the metabolic profiles of these three major types of nephropathies and identify potential metabolic biomarkers.
Material and methods:
Kidney samples of 20 sex- and age-matched patients with biopsy-proven IgAN, MN, DN, and controls without any kidney diseases were included. Ultra-high-performance liquid chromatography–mass spectrometry analysis was conducted. The t-test was used to evaluate the statistical significance of the identified metabolites. Metabolic pathways were analyzed using the Kyoto Encyclopedia of Genes and Genomes (KEGG). Specificity, sensitivity, and area under the curve (AUC) were calculated to evaluate the predictive performance of metabolites.
Results:
Among 557 identified differential metabolites, only 118 were found in all three comparison groups. Differential metabolites of IgAN vs. controls were significantly enriched in arachidonic acid metabolism, starch and sucrose metabolism, ferroptosis, and other pathways. In the DN group, metabolites were mainly enriched in phenylalanine, tyrosine and tryptophan biosynthesis, histidine metabolism, etc. MN-enriched pathways included steroid hormone biosynthesis, neuroactive ligand-receptor interaction, and bile secretion. In the positive mode, cumulative AUC values for comparison pairs IgAN vs. controls, MN vs controls, and DN vs controls were 0.965, 0.972, and 0.573, respectively, whereas in the negative mode the AUC values of all three pairs were slightly above 0.65.
Conclusions:
IgAN, MN, and DN have similar but distinct metabolic profiles. Only positive node metabolites of IgAN and MN exhibited high predictive performance.
REFERENCES (33)
1.
Vahedian-Azimi A, Beni FH, Fras Z, et al. Effects of statins on the incidence and outcomes of acute kidney injury in critically ill patients: a systematic review and meta-analysis. Arch Med Sci 2023; 19: 952-64.
2.
Katsiki N, Kolovou G, Melidonis A, Banach M. The Cardiac- Kidney-Liver (CKL) syndrome: the “real entity” of type 2 diabetes mellitus. Arch Med Sci 2024; 20: 207-15.
3.
Guo X, Tie X, Zhang Y, et al. Management and clinical outcomes of membranous nephropathy, IgA nephropathy, and minimal change disease two years post-kidney biopsy. Kidney Blood Press Res 2024; 49: 345-54.
4.
Stamellou E, Seikrit C, Tang SCW, et al. IgA nephropathy. Nat Rev Dis Primers 2023; 9: 67.
5.
Nieto-Gañán I, Iturrieta-Zuazo I, Rita C, Carrasco-Sayale- ro Á. Revisiting immunological and clinical aspects of membranous nephropathy. Clin Immunol 2022; 237: 108976.
6.
Fox CS, Matsushita K, Woodward M, et al. Associations of kidney disease measures with mortality and end-stage renal disease in individuals with and without diabetes: a meta-analysis. Lancet 2012; 380: 1662-73.
7.
Furlani IL, da Cruz Nunes E, Canuto GAB, Macedo AN, Oliveira RV. Liquid chromatography-mass spectrometry for clinical metabolomics: an overview. Adv Exp Med Biol 2021; 1336: 179-213.
8.
Pereira PR, Carrageta DF, Oliveira PF, Rodrigues A, Alves MG, Monteiro MP. Metabolomics as a tool for the early diagnosis and prognosis of diabetic kidney disease. Med Res Rev 2022; 42: 1518-44.
9.
Wang H, Ainiwaer A, Song Y, et al. Perturbed gut microbiome and fecal and serum metabolomes are associated with chronic kidney disease severity. Microbiome 2023; 11: 3.
10.
Harvey FC, Collao V, Bhattacharya SK. High-resolution liquid chromatography-mass spectrometry for lipidomics. Methods Mol Biol 2023; 2625: 57-63.
11.
González-Domínguez Á, Armeni M, Savolainen O, Lechuga- Sancho AM, Landberg R, González-Domínguez R. Untargeted metabolomics based on liquid chromatography-mass spectrometry for the analysis of plasma and erythrocyte samples in childhood obesity. Methods Mol Biol 2023; 2571: 115-22.
12.
Saucedo AL, Perales-Quintana MM, Paniagua-Vega D, Sanchez-Martinez C, Cordero-Perez P, Minsky NW. Chronic kidney disease and the search for new biomarkers for early diagnosis. Curr Med Chem 2018; 25: 3719-47.
13.
Li H, Li D, Ledru N, et al. Transcriptomic, epigenomic, and spatial metabolomic cell profiling redefines regional human kidney anatomy. Cell Metab 2024; 36: 1105-25.e10.
14.
Lecamwasam A, Mansell T, Ekinci EI, Saffery R, Dwyer KM. Blood plasma metabolites in diabetes-associated chronic kidney disease: a focus on lipid profiles and cardiovascular risk. Front Nutr 2022; 9: 821209.
15.
Fromentin S, Forslund SK, Chechi K, et al. Microbiome and metabolome features of the cardiometabolic disease spectrum. Nat Med 2022; 28: 303-14.
16.
Uwaezuoke SN, Muoneke UV, Mbanefo NR. The supportive treatment of IgA nephropathy and idiopathic nephrotic syndrome: how useful are omega-3 polyunsaturated fatty acids? Int J Nephrol Renovasc Dis 2020; 13: 27-35.
17.
Wu H, Tang D, Yun M, et al. Metabolic dysfunctions of intestinal fatty acids and tryptophan reveal immuno- inflammatory response activation in IgA nephropathy. Front Med 2022; 9: 811526.
18.
Cui C, Wang C, Han S, Yu D, Zhu L, Jiang P. Impact of a long-term high-fructose diet on systemic metabolic profiles of mice. FASEB Bioadv 2022; 4: 560-72.
19.
Wang X, He Q, Chen Q, et al. Network pharmacology combined with metabolomics to study the mechanism of Shenyan Kangfu Tablets in the treatment of diabetic nephropathy. J Ethnopharmacol 2021; 270: 113817.
20.
Zhang S, Li X, Luo H, Fang ZZ, Ai H. Role of aromatic amino acids in pathogeneses of diabetic nephropathy in Chinese patients with type 2 diabetes. J Diabetes Complications 2020; 34: 107667.
21.
Luo HH, Li J, Feng XF, et al. Plasma phenylalanine and tyrosine and their interactions with diabetic nephropathy for risk of diabetic retinopathy in type 2 diabetes. BMJ Open Diabetes Res Care 2020; 8: e000877.
22.
Liu Y, Chen X, Liu Y, et al. Metabolomic study of the protective effect of Gandi capsule for diabetic nephropathy. Chem Biol Interact 2019; 314: 108815.
23.
Zhao J, He K, Du H, et al. Bioinformatics prediction and experimental verification of key biomarkers for diabetic kidney disease based on transcriptome sequencing in mice. PeerJ 2022; 10: e13932.
24.
Li J, Cao Y, Lu R, et al. Integrated fecal microbiome and serum metabolomics analysis reveals abnormal changes in rats with immunoglobulin a nephropathy and the intervention effect of Zhen Wu Tang. Front Pharmacol 2020; 11: 606689.
25.
Xue R, Wang Y, Geng L, et al. Comprehensive analysis of the gene expression profile of the male and female BTBR mice with diabetic nephropathy. Int J Biol Macromol 2024; 257: 128720.
26.
Ren H, Lv W, Shang Z, et al. Identifying functional subtypes of IgA nephropathy based on three machine learning algorithms and WGCNA. BMC Med Genomics 2024; 17: 61.
27.
Yang J, Pontoglio M, Terzi F. Bile acids and Farnesoid X Receptor in renal pathophysiology. Nephron 2024; 148: 618-30.
28.
Niessen WM. Fragmentation of toxicologically relevant drugs in positive-ion liquid chromatography-tandem mass spectrometry. Mass Spectrom Rev 2011; 30: 626-63.
29.
Niessen WM. Fragmentation of toxicologically relevant drugs in negative-ion liquid chromatography-tandem mass spectrometry. Mass Spectrom Rev 2012; 31: 626-65.
30.
Ye M, Tang D, Li W, et al. Serum metabolomics analysis reveals metabolite profile and key biomarkers of idiopathic membranous nephropathy. PeerJ 2023; 11: e15167.
31.
Dong L, Tan J, Zhong Z, Tang Y, Qin W. Altered serum metabolic profile in patients with IgA nephropathy. Clin Chim Acta 2023; 549: 117561.
32.
Lin HT, Cheng ML, Lo CJ, Lin G, Liu FC. Metabolomic signature of diabetic kidney disease in cerebrospinal fluid and plasma of patients with type 2 diabetes using liquid chromatography-mass spectrometry. Diagnostics 2022; 12: 2626.
33.
Liu S, Gui Y, Wang MS, et al. Serum integrative omics reveals the landscape of human diabetic kidney disease. Mol Metab 2021; 54: 101367.
Share
RELATED ARTICLE
We process personal data collected when visiting the website. The function of obtaining information about users and their behavior is carried out by voluntarily entered information in forms and saving cookies in end devices. Data, including cookies, are used to provide services, improve the user experience and to analyze the traffic in accordance with the Privacy policy. Data are also collected and processed by Google Analytics tool (more).
You can change cookies settings in your browser. Restricted use of cookies in the browser configuration may affect some functionalities of the website.
You can change cookies settings in your browser. Restricted use of cookies in the browser configuration may affect some functionalities of the website.